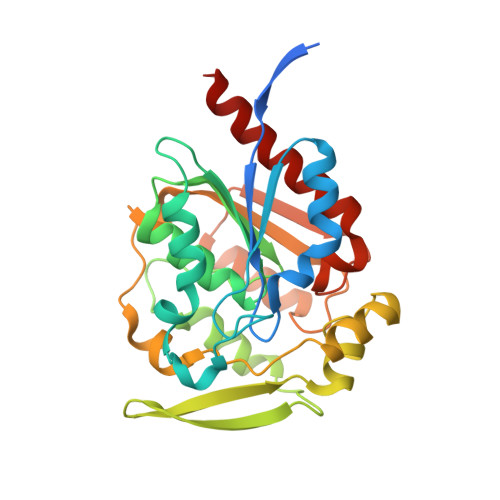
Structural and functional analysis of hydroxynitrile lyase from Baliospermum montanum with crystal structure, molecular dynamics and enzyme kinetics
Nakano, S., Dadashipour, M., Asano, Y.(2014) Biochim Biophys Acta 1844: 2059-2067
- PubMed: 25220808 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2014.09.004
- Primary Citation Related Structures:
3WWO, 3WWP - PubMed Abstract:
Hydroxynitrile lyases (HNLs) catalyze degradation of cyanohydrins to hydrogen cyanide and the corresponding ketone or aldehyde. HNLs can also catalyze the reverse reaction, i.e., synthesis of cyanohydrins. Although several crystal structures of S-selective hydroxynitrile lyases (S-HNLs) have been reported, it remains unknown whether and how dynamics at the active site of S-HNLs influence their broad substrate specificity and affinity. In this study, we analyzed the structure, dynamics and function of S-HNL from Baliospermum montanum (BmHNL), which has an α/β hydrolase fold. Two crystal structures of BmHNL, apo1 and apo2, were determined at 2.55 and 1.9Å, respectively. Structural comparison between BmHNL (apo2) and S-HNL from Hevea brasiliensis with (S)-mandelonitrile bound to the active site revealed that hydrophobic residues at the entrance region of BmHNL formed hydrophobic interactions with the benzene ring of the substrate. The flexible structures of these hydrophobic residues were confirmed by a 15ns molecular dynamics simulation. This flexibility regulated the size of the active site cavity, enabling binding of various substrates to BmHNL. The high affinity of BmHNL toward substrates containing a benzene ring was also confirmed by comparing the kinetics of BmHNL and S-HNL from Manihot esculenta. Taken together, the results indicated that the flexibility and placement of the residues are important for the broad substrate specificity of S-HNLs.
- Biotechnology Research Center and Department of Biotechnology, Toyama Prefectural University, 5180 Kurokawa, Imizu, Toyama 939-0398, Japan; Asano Active Enzyme Molecule Project, ERATO, JST, 5180 Kurokawa, Imizu, Toyama 939-0398, Japan.
Organizational Affiliation: