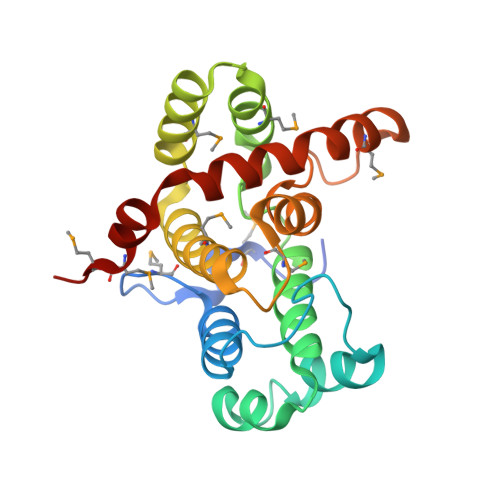
Structural basis for the effects of Ser387 phosphorylation of MgcRacGAP on its GTPase-activating activities for CDC42 and RHOA.
Murayama, K., Kato-Murayama, M., Hosaka, T., Kitamura, T., Yokoyama, S., Shirouzu, M.(2024) J Struct Biol 216: 108151-108151
- PubMed: 39522789
- DOI: https://doi.org/10.1016/j.jsb.2024.108151
- Primary Citation of Related Structures:
3WPQ, 3WPS, 5C2J, 5C2K - PubMed Abstract:
MgcRacGAP is a GTPase-activating protein (GAP) for the Rho family GTPases. During cytokinesis, MgcRacGAP localizes to the midbody, where it activates the GTPase activity of Rho family GTPases to facilitate cytokinesis. In the midbody, Aurora B phosphorylates Ser387 within the GAP domain of human MgcRacGAP, a modification that is suggested to influence GTPase preference. However, there are conflicting reports, with some studies indicating that Ser387 phosphorylation does not alter the GTPase preference of MgcRacGAP. This controversy highlights the need for a deeper understanding of the molecular interactions involved, which can be clarified through structural analyses. In the present study, we determined the crystal structures of the wild-type MgcRacGAP GAP domain complexed with CDC42•GDP•AlF 4 - and the S378D phosphomimetic mutant GAP domain fused with RHOA•GDP•AlF 4 - . Additionally, crystal structures of the GAP domains were determined for the S387D and S387A mutants. Our analysis revealed that neither GTPase binding nor S387D mutation affected the overall structure of the GAP domain. However, comparison of the CDC42•MgcRacGAP (wild-type) complex with the RHOA-MgcRacGAP(S378D) fusion protein structure indicated that the S387D mutation caused positional shifts in both CDC42 and RHOA relative to MgcRacGAP. These shifts reduced interactions with CDC42 more severely than those with RHOA. In fact, the S387D mutation decreased the GTPase-activating activity of MgcRacGAP toward CDC42, while its impact on RHOA was only moderate. This difference in the rate of activity reduction may play an important role in GTPase preference.
- Division of Biomedical Measurements and Diagnostics, Graduate School of Biomedical Engineering, Tohoku University, 2-1 Seiryo, Aoba, Sendai 980-8575, Japan; RIKEN Systems and Structural Biology Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan; Laboratory for Protein Functional and Structural Biology, RIKEN Center for Biosystems Dynamics Research, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: