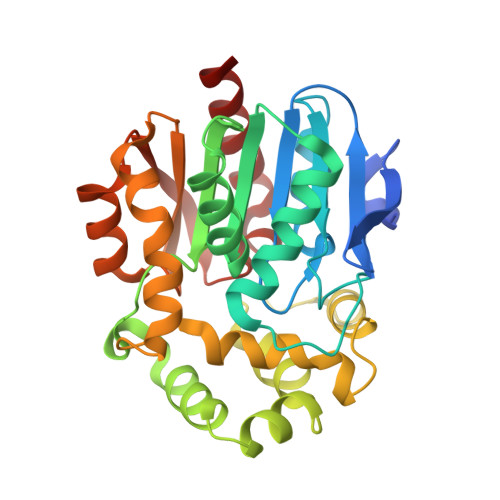
Crystal structure of the novel haloalkane dehalogenase DatA from Agrobacterium tumefaciens C58 reveals a special halide-stabilizing pair and enantioselectivity mechanism.
Guan, L., Yabuki, H., Okai, M., Ohtsuka, J., Tanokura, M.(2014) Appl Microbiol Biotechnol 98: 8573-8582
- PubMed: 24770384 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-014-5751-2
- Primary Citation Related Structures:
3WI7, 3WIB - PubMed Abstract:
A novel haloalkane dehalogenase DatA from Agrobacterium tumefaciens C58 belongs to the HLD-II subfamily and hydrolyzes brominated and iodinated compounds, leading to the generation of the corresponding alcohol, a halide ion, and a proton. Because DatA possesses a unique Asn-Tyr pair instead of the Asn-Trp pair conserved among the subfamily members, which was proposed to keep the released halide ion stable, the structural basis for its reaction mechanism should be elucidated. Here, we determined the crystal structures of DatA and its Y109W mutant at 1.70 and 1.95 Å, respectively, and confirmed the location of the active site by using its novel competitive inhibitor. The structural information from these two crystal structures and the docking simulation suggested that (i) the replacement of the Asn-Tyr pair with the Asn-Trp pair increases the binding affinity for some halogenated compounds, such as 1,3-dibromopropane, mainly due to the electrostatic interaction between Trp109 and halogenated compounds and the change of substrate-binding mode caused by the interaction and (ii) the primary halide-stabilizing residue is only Asn43 in the wild-type DatA, while Tyr109 is a secondary halide-stabilizing residue. Furthermore, docking simulation using the crystal structures of DatA indicated that its enantioselectivity is determined by the large and small spaces around the halogen-binding site.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo, Tokyo, 113-8657, Japan.
Organizational Affiliation: