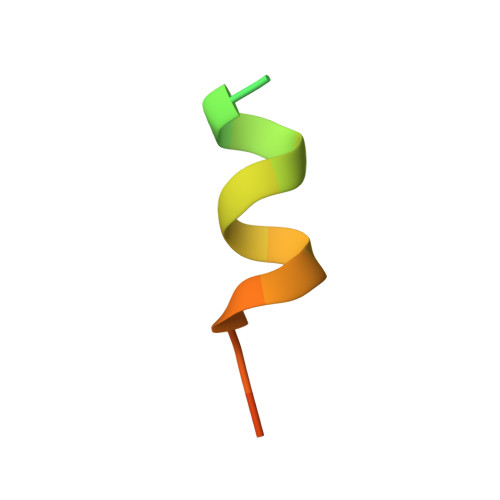
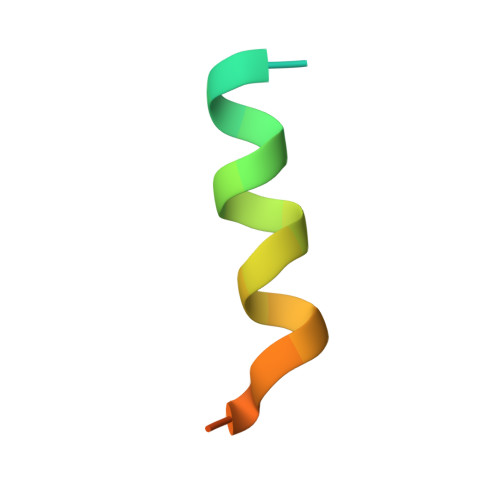
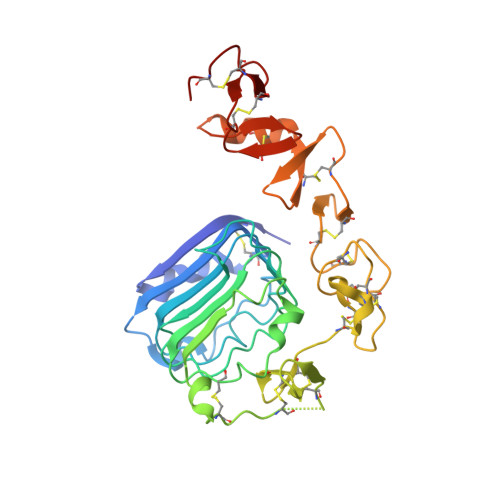
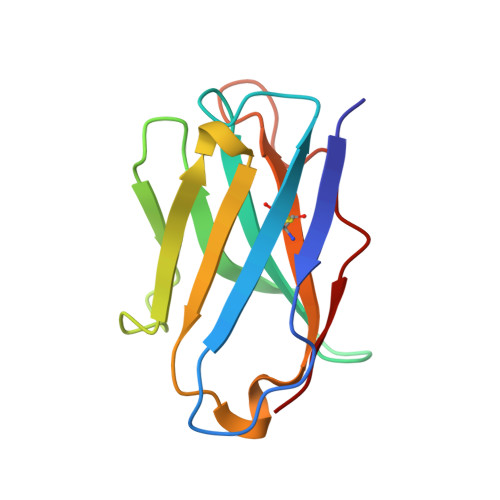
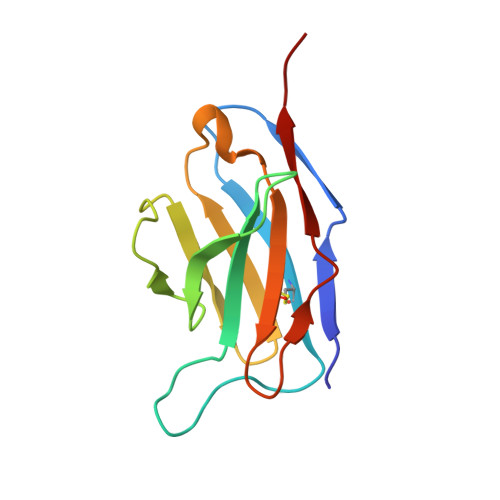
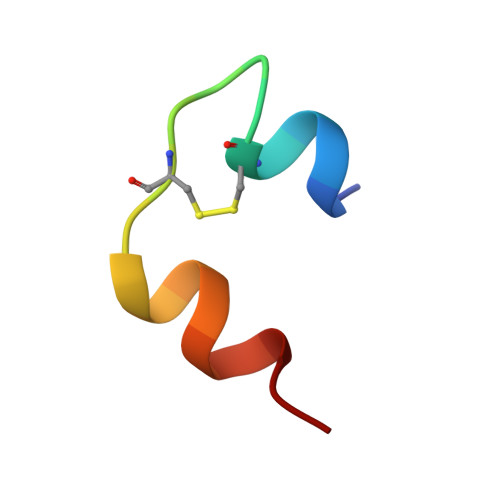
How insulin engages its primary binding site on the insulin receptor
Menting, J.G., Whittaker, J., Margetts, M.B., Whittaker, L.J., Kong, G.K.-W., Smith, B.J., Watson, C.J., Zakova, L., Kletvikova, E., Jiracek, J., Chan, S.J., Steiner, D.F., Dodson, G.G., Brzozowski, A.M., Weiss, M.A., Ward, C.W., Lawrence, M.C.(2013) Nature 493: 241-245
- PubMed: 23302862 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature11781
- Primary Citation Related Structures:
3W11, 3W12, 3W13, 5KQV - PubMed Abstract:
Insulin receptor signalling has a central role in mammalian biology, regulating cellular metabolism, growth, division, differentiation and survival. Insulin resistance contributes to the pathogenesis of type 2 diabetes mellitus and the onset of Alzheimer's disease; aberrant signalling occurs in diverse cancers, exacerbated by cross-talk with the homologous type 1 insulin-like growth factor receptor (IGF1R). Despite more than three decades of investigation, the three-dimensional structure of the insulin-insulin receptor complex has proved elusive, confounded by the complexity of producing the receptor protein. Here we present the first view, to our knowledge, of the interaction of insulin with its primary binding site on the insulin receptor, on the basis of four crystal structures of insulin bound to truncated insulin receptor constructs. The direct interaction of insulin with the first leucine-rich-repeat domain (L1) of insulin receptor is seen to be sparse, the hormone instead engaging the insulin receptor carboxy-terminal α-chain (αCT) segment, which is itself remodelled on the face of L1 upon insulin binding. Contact between insulin and L1 is restricted to insulin B-chain residues. The αCT segment displaces the B-chain C-terminal β-strand away from the hormone core, revealing the mechanism of a long-proposed conformational switch in insulin upon receptor engagement. This mode of hormone-receptor recognition is novel within the broader family of receptor tyrosine kinases. We support these findings by photo-crosslinking data that place the suggested interactions into the context of the holoreceptor and by isothermal titration calorimetry data that dissect the hormone-insulin receptor interface. Together, our findings provide an explanation for a wealth of biochemical data from the insulin receptor and IGF1R systems relevant to the design of therapeutic insulin analogues.
- Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Parkville, Victoria 3052, Australia.
Organizational Affiliation: