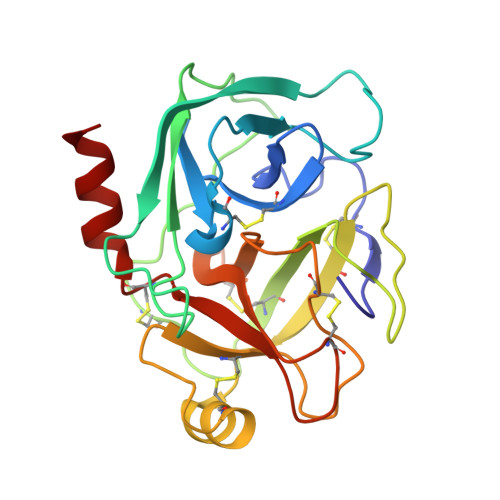
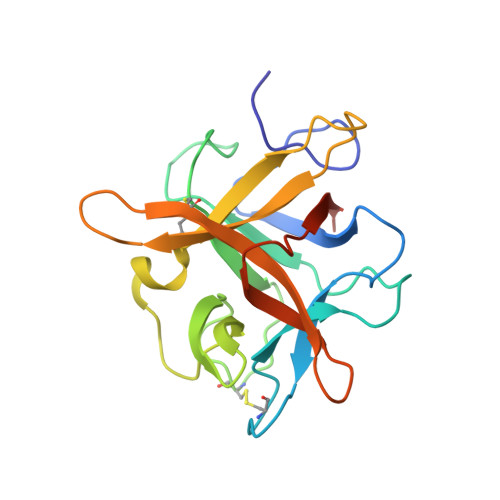
Role of remote scaffolding residues in the inhibitory loop pre-organization, flexibility, rigidification and enzyme inhibition of serine protease inhibitors
Majumder, S., Khamrui, S., Dasgupta, J., Dattagupta, J.K., Sen, U.(2012) Biochim Biophys Acta 1824: 882-890
- PubMed: 22709512 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2012.04.009
- Primary Citation Related Structures:
3I29, 3QYD, 3VEQ - PubMed Abstract:
Canonical serine protease inhibitors interact with cognate enzymes through the P3-P2' region of the inhibitory loop while its scaffold hardly makes any contact. Neighboring scaffolding residues like Arginines or Asparagine shape-up the inhibitory loop and favor the resynthesis of cleaved scissile bond. However, role of remote scaffolding residues, which are not involved in religation, was not properly explored. Crystal structures of two engineered winged bean chymotrypsin inhibitor (WCI) complexed with Bovine trypsin (BPT) namely L65R-WCI:BPT and F64Y/L65R-WCI:BPT show that the inhibitory loop of these engineered inhibitors are recognized and rigidified properly at the enzyme active site like other strong trypsin inhibitors. Chimeric protein ETI(L)-WCI(S), having a loop of Erythrina caffra Trypsin Inhibitor, ETI on the scaffold of WCI, was previously shown to behave like substrate. Non-canonical structure of the inhibitory loop and its flexibility are attributed to the presence of smaller scaffolding residues which cannot act as barrier to the inhibitory loop like in ETI. Double mutant A76R/L115Y-(ETI(L)-WCI(S)), where the barrier is reintroduced on ETI(L)-WCI(S), shows regaining of inhibitory activity. The structure of A76R/L115Y-(ETI(L)-WCI(S)) along with L65R-WCI:BPT and F64Y/L65R-WCI:BPT demonstrate here that the lost canonical conformation of the inhibitory loop is fully restored and loop flexibility is dramatically reduced. Therefore, residues at the inhibitory loop interact with the enzyme playing the primary role in recognition and binding but scaffolding residues having no direct interaction with the enzyme are crucial for rigidification event and the inhibitory potency. B-factor analysis indicates that the amount of inhibitory loop rigidification varies between different inhibitor families.
- Crystallography and Molecular Biology Division, Saha Institute of Nuclear Physics, Kolkata, India.
Organizational Affiliation: