Teaching an old scaffold new tricks: monobodies constructed using alternative surfaces of the FN3 scaffold.
Koide, A., Wojcik, J., Gilbreth, R.N., Hoey, R.J., Koide, S.(2012) J Mol Biology 415: 393-405
- PubMed: 22198408 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.12.019
- Primary Citation Related Structures:
3RZW, 3UYO - PubMed Abstract:
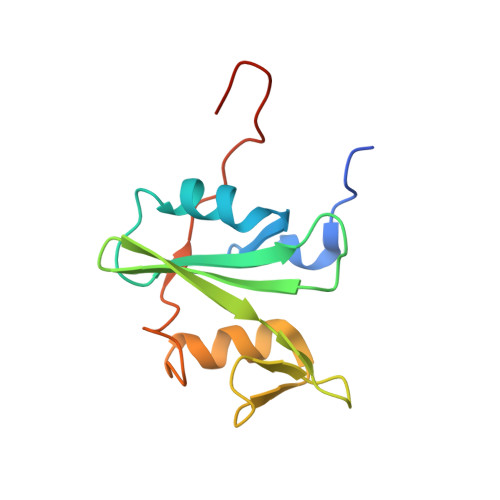
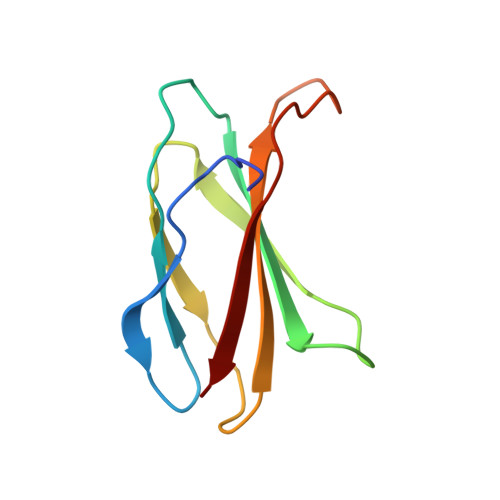
The fibronectin type III domain (FN3) has become one of the most widely used non-antibody scaffolds for generating new binding proteins. Because of its structural homology to the immunoglobulin domain, combinatorial libraries of FN3 designed to date have primarily focused on introducing amino acid diversity into three loops that are equivalent to antibody complementarity-determining regions. Here, we report an FN3 library that utilizes alternative positions for presenting amino acid diversity. We diversified positions on a β-sheet and surface loops that together form a concave surface. The new library produced binding proteins (termed "monobodies") to multiple target proteins, generally with similar efficacy as the original, loop-focused library. The crystal structure of a monobody generated from the new library in complex with its target, the Abl SH2 domain, revealed that a concave surface of the monobody, as intended in our design, bound to a convex surface of the target with the interface area being among the largest of published structures of monobody-target complexes. This mode of interaction differs from a common binding mode for single-domain antibodies and antibody mimics in which recognition loops recognize clefts in targets. Together, this work illustrates the utilization of different surfaces of a single immunoglobulin-like scaffold to generate binding proteins with distinct characteristics.
- Department of Biochemistry and Molecular Biology, The University of Chicago, 929 East 57th Street, Chicago,IL 60637, USA.
Organizational Affiliation: