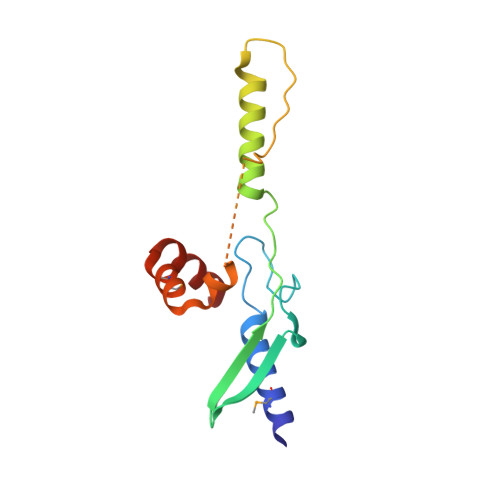
The mammalian DUF59 protein Fam96a forms two distinct types of domain-swapped dimer.
Chen, K.E., Richards, A.A., Ariffin, J.K., Ross, I.L., Sweet, M.J., Kellie, S., Kobe, B., Martin, J.L.(2012) Acta Crystallogr D Biol Crystallogr 68: 637-648
- PubMed: 22683786 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912006592
- Primary Citation Related Structures:
3UX2, 3UX3 - PubMed Abstract:
Fam96a mRNA, which encodes a mammalian DUF59 protein, is enriched in macrophages. Recombinant human Fam96a forms stable monomers and dimers in solution. Crystal structures of these two forms revealed that each adopts a distinct type of domain-swapped dimer, one of which is stabilized by zinc binding. Two hinge loops control Fam96a domain swapping; both are flexible and highly conserved, suggesting that domain swapping may be a common feature of eukaryotic but not bacterial DUF59 proteins. The derived monomer fold of Fam96a diverges from that of bacterial DUF59 counterparts in that the C-terminal region of Fam96a is much longer and is positioned on the opposite side of the N-terminal core fold. The putative metal-binding site of bacterial DUF59 proteins is not conserved in Fam96a, but Fam96a interacts tightly in vitro with Ciao1, the cytosolic iron-assembly protein. Moreover, Fam96a and Ciao1 can be co-immunoprecipitated, suggesting that the interaction also occurs in vivo. Although predicted to have a signal peptide, it is shown that Fam96a is cytoplasmic. The data reveal that eukaryotic DUF59 proteins share intriguing characteristics with amyloidogenic proteins.
- Institute for Molecular Bioscience, Division of Chemistry and Structural Biology, University of Queensland, Brisbane, Queensland 4072, Australia.
Organizational Affiliation: