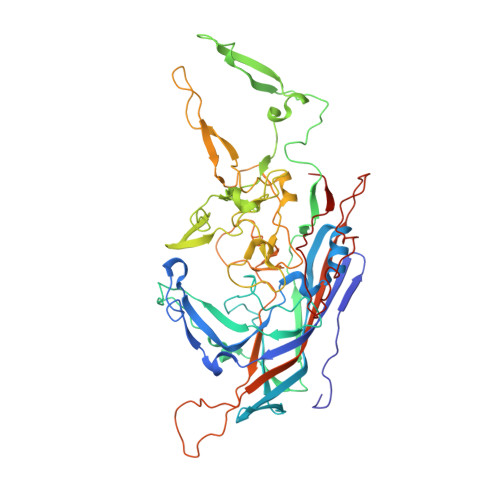
Structural insight into the unique properties of adeno-associated virus serotype 9.
Dimattia, M.A., Nam, H.J., Van Vliet, K., Mitchell, M., Bennett, A., Gurda, B.L., McKenna, R., Olson, N.H., Sinkovits, R.S., Potter, M., Byrne, B.J., Aslanidi, G., Zolotukhin, S., Muzyczka, N., Baker, T.S., Agbandje-McKenna, M.(2012) J Virol 86: 6947-6958
- PubMed: 22496238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.07232-11
- Primary Citation Related Structures:
3UX1 - PubMed Abstract:
Adeno-associated virus serotype 9 (AAV9) has enhanced capsid-associated tropism for cardiac muscle and the ability to cross the blood-brain barrier compared to other AAV serotypes. To help identify the structural features facilitating these properties, we have used cryo-electron microscopy (cryo-EM) and three-dimensional image reconstruction (cryo-reconstruction) and X-ray crystallography to determine the structure of the AAV9 capsid at 9.7- and 2.8-Å resolutions, respectively. The AAV9 capsid exhibits the surface topology conserved in all AAVs: depressions at each icosahedral two-fold symmetry axis and surrounding each five-fold axis, three separate protrusions surrounding each three-fold axis, and a channel at each five-fold axis. The AAV9 viral protein (VP) has a conserved core structure, consisting of an eight-stranded, β-barrel motif and the αA helix, which are present in all parvovirus structures. The AAV9 VP differs in nine variable surface regions (VR-I to -IX) compared to AAV4, but at only three (VR-I, VR-II, and VR-IV) compared to AAV2 and AAV8. VR-I differences modify the raised region of the capsid surface between the two-fold and five-fold depressions. The VR-IV difference produces smaller three-fold protrusions in AAV9 that are less "pointed" than AAV2 and AAV8. Significantly, residues in the AAV9 VRs have been identified as important determinants of cellular tropism and transduction and dictate its antigenic diversity from AAV2. Hence, the AAV9 VRs likely confer the unique infection phenotypes of this serotype.
- Department of Biochemistry and Molecular Biology, Center for Structural Biology, McKnight Brain Institute, College of Medicine, University of Florida, Gainesville, Florida, USA.
Organizational Affiliation: