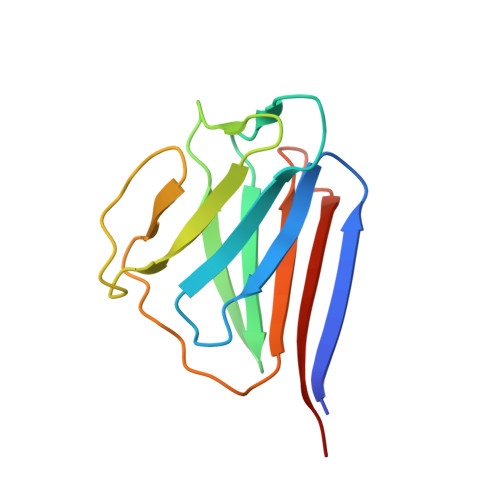
Crystal structures of the coil 2B fragment and the globular tail domain of human lamin B1.
Ruan, J., Xu, C., Bian, C., Lam, R., Wang, J.P., Kania, J., Min, J., Zang, J.(2012) FEBS Lett 586: 314-318
- PubMed: 22265972 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2012.01.007
- Primary Citation Related Structures:
3TYY, 3UMN - PubMed Abstract:
We present here the crystal structures of human lamin B1 globular tail domain and coiled 2B domain, which adopt similar folds to Ig-like domain and coiled-coil domain of lamin A, respectively. Despite the overall similarity, we found an extra intermolecular disulfide bond in the lamin B1 coil 2B domain, which does not exist in lamin A/C. In addition, the structural analysis indicates that interactions at the lamin B1 homodimer interface are quite different from those of lamin A/C. Thus our research not only reveals the diversely formed homodimers among lamin family members, but also sheds light on understanding the important roles of lamin B1 in forming the nuclear lamina matrix.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, People's Republic of China.
Organizational Affiliation: