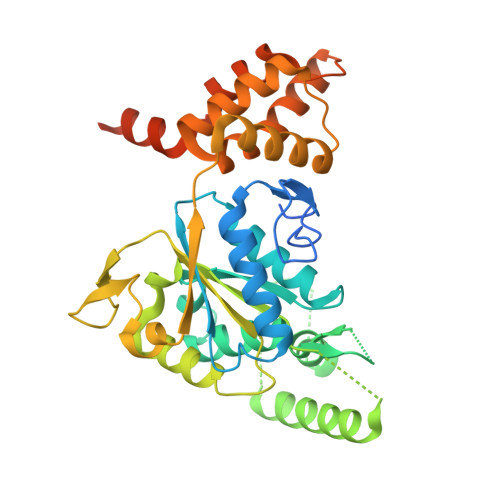
Large-Scale Conformational Flexibility Determines the Properties of AAA+ TIP49 ATPases.
Petukhov, M., Dagkessamanskaja, A., Bommer, M., Barrett, T., Tsaneva, I., Yakimov, A., Queval, R., Shvetsov, A., Khodorkovskiy, M., Kas, E., Grigoriev, M.(2012) Structure 20: 1321-1331
- PubMed: 22748767 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.05.012
- Primary Citation Related Structures:
3UK6 - PubMed Abstract:
The TIP49a and TIP49b proteins belong to the family of AAA+ ATPases and play essential roles in vital processes such as transcription, DNA repair, snoRNP biogenesis, and chromatin remodeling. We report the crystal structure of a TIP49b hexamer and the comparative analysis of large-scale conformational flexibility of TIP49a, TIP49b, and TIP49a/TIP49b complexes using molecular modeling and molecular dynamics simulations in a water environment. Our results establish key principles of domain mobility that affect protein conformation and biochemical properties, including a mechanistic basis for the downregulation of ATPase activity upon protein hexamerization. These approaches, applied to the lik-TIP49b mutant reported to possess enhanced DNA-independent ATPase activity, help explain how a three-amino acid insertion remotely affects the structure and conformational dynamics of the ATP binding and hydrolysis pocket while uncoupling ATP hydrolysis from DNA binding. This might be similar to the effects of conformations adopted by TIP49 heterohexamers.
- Department of Biophysics, Saint Petersburg State Polytechnical University, Saint Petersburg 197376, Russia.
Organizational Affiliation: