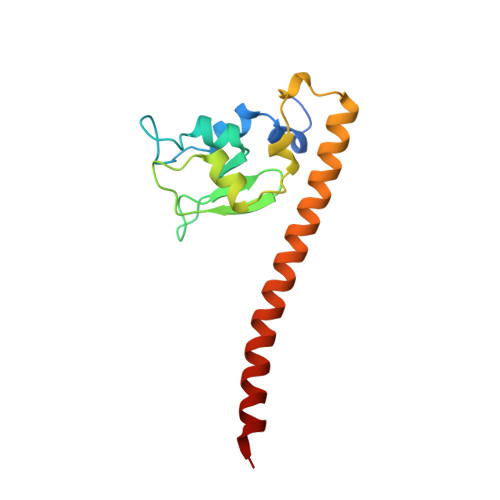
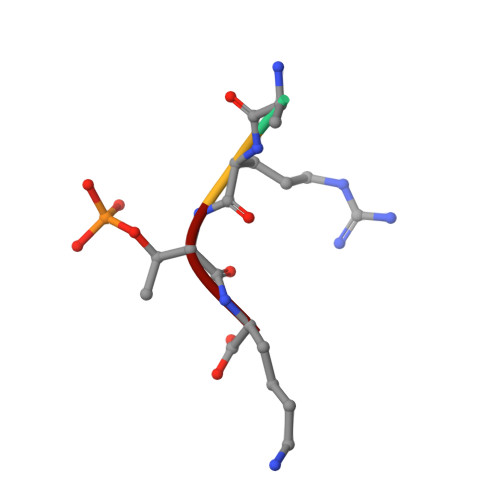
Molecular basis for phosphospecific recognition of histone H3 tails by Survivin paralogues at inner centromeres.
Niedzialkowska, E., Wang, F., Porebski, P.J., Minor, W., Higgins, J.M., Stukenberg, P.T.(2012) Mol Biol Cell 23: 1457-1466
- PubMed: 22357620 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.E11-11-0904
- Primary Citation Related Structures:
3UEC, 3UED, 3UEE, 3UEF, 3UEG, 3UEH, 3UEI - PubMed Abstract:
Survivin, a subunit of the chromosome passenger complex (CPC), binds the N-terminal tail of histone H3, which is phosphorylated on T3 by Haspin kinase, and localizes the complex to the inner centromeres. We used x-ray crystallography to determine the residues of Survivin that are important in binding phosphomodified histone H3. Mutation of amino acids that interact with the histone N-terminus lowered in vitro tail binding affinity and reduced CPC recruitment to the inner centromere in cells, validating our solved structures. Phylogenetic analysis shows that nonmammalian vertebrates have two Survivin paralogues, which we name class A and B. A distinguishing feature of these paralogues is an H-to-R change in an amino acid that interacts with the histone T3 phosphate. The binding to histone tails of the human class A paralogue, which has a histidine at this position, is sensitive to changes around physiological pH, whereas Xenopus Survivin class B is less so. Our data demonstrate that Survivin paralogues have different characteristics of phosphospecific binding to threonine-3 of histone H3, providing new insight into the biology of the inner centromere.
- Department of Biochemistry and Molecular Genetics, University of Virginia School of Medicine, Charlottesville, VA 22908, USA.
Organizational Affiliation: