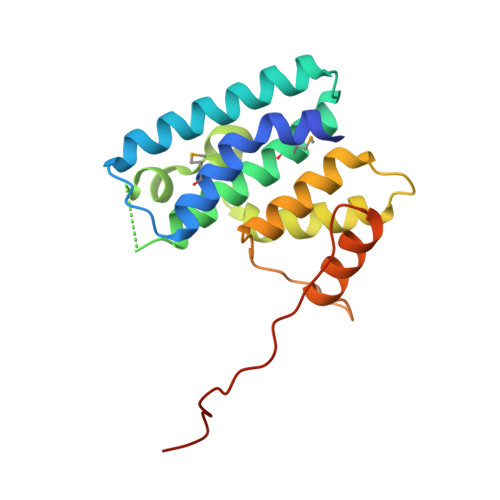
Structural characterization and oligomerization of the TssL protein, a component shared by bacterial type VI and type IVb secretion systems.
Durand, E., Zoued, A., Spinelli, S., Watson, P.J., Aschtgen, M.S., Journet, L., Cambillau, C., Cascales, E.(2012) J Biological Chem 287: 14157-14168
- PubMed: 22371492 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.338731
- Primary Citation Related Structures:
3U66 - PubMed Abstract:
The Type VI secretion system (T6SS) is a macromolecular system distributed in Gram-negative bacteria, responsible for the secretion of effector proteins into target cells. The T6SS has a broad versatility as it can target both eukaryotic and prokaryotic cells. It is therefore involved in host pathogenesis or killing neighboring bacterial cells to colonize a new niche. At the architecture level, the T6SS core apparatus is composed of 13 proteins, which assemble in two subcomplexes. One of these subcomplexes, composed of subunits that share structural similarities with bacteriophage tail and baseplate components, is anchored to the cell envelope by the membrane subcomplex. This latter is constituted of at least three proteins, TssL, TssM, and TssJ. The crystal structure of the TssJ outer membrane lipoprotein and its interaction with the inner membrane TssM protein have been recently reported. TssL and TssM share sequence homology and characteristics with two components of the Type IVb secretion system (T4bSS), IcmH/DotU and IcmF, respectively. In this study, we report the crystal structure of the cytoplasmic domain of the TssL inner membrane protein from the enteroaggregative Escherichia coli Sci-1 T6SS. It folds as a hook-like structure composed of two three-helix bundles. Two TssL molecules associate to form a functional complex. Although the TssL trans-membrane segment is the main determinant of self-interaction, contacts between the cytoplasmic domains are required for TssL function. Based on sequence homology and secondary structure prediction, we propose that the TssL structure is the prototype for the members of the TssL and IcmH/DotU families.
- Aix-Marseille Université, Architecture et Fonction des Macromolécules Biologiques, Campus de Luminy, Case 932, 13288 Marseille Cedex 09, France.
Organizational Affiliation: