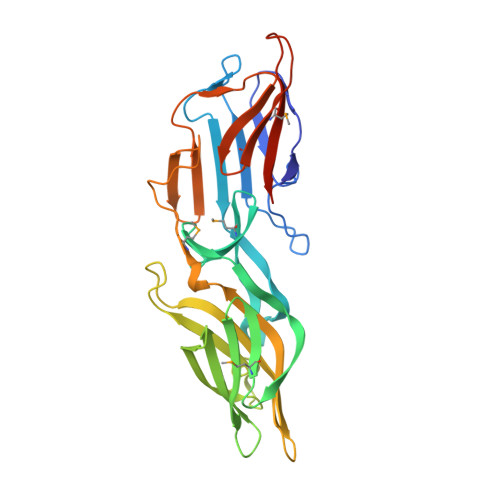
Structure of the surface layer of the methanogenic archaean Methanosarcina acetivorans.
Arbing, M.A., Chan, S., Shin, A., Phan, T., Ahn, C.J., Rohlin, L., Gunsalus, R.P.(2012) Proc Natl Acad Sci U S A 109: 11812-11817
- PubMed: 22753492 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1120595109
- Primary Citation Related Structures:
3U2G, 3U2H - PubMed Abstract:
Archaea have a self-assembling proteinaceous surface (S-) layer as the primary and outermost boundary of their cell envelopes. The S-layer maintains structural rigidity, protects the organism from adverse environmental elements, and yet provides access to all essential nutrients. We have determined the crystal structure of one of the two "homologous" tandem polypeptide repeats that comprise the Methanosarcina acetivorans S-layer protein and propose a high-resolution model for a microbial S-layer. The molecular features of our hexameric S-layer model recapitulate those visualized by medium resolution electron microscopy studies of microbial S-layers and greatly expand our molecular view of S-layer dimensions, porosity, and symmetry. The S-layer model reveals a negatively charged molecular sieve that presents both a charge and size barrier to restrict access to the cell periplasmic-like space. The β-sandwich folds of the S-layer protein are structurally homologous to eukaryotic virus envelope proteins, suggesting that Archaea and viruses have arrived at a common solution for protective envelope structures. These results provide insight into the evolutionary origins of primitive cell envelope structures, of which the S-layer is considered to be among the most primitive: it also provides a platform for the development of self-assembling nanomaterials with diverse functional and structural properties.
- University of California at Los Angeles-US Department of Energy (UCLA-DOE) Institute for Genomics and Proteomics, University of California, Los Angeles, CA 90095, USA. marbing@mbi.ucla.edu
Organizational Affiliation: