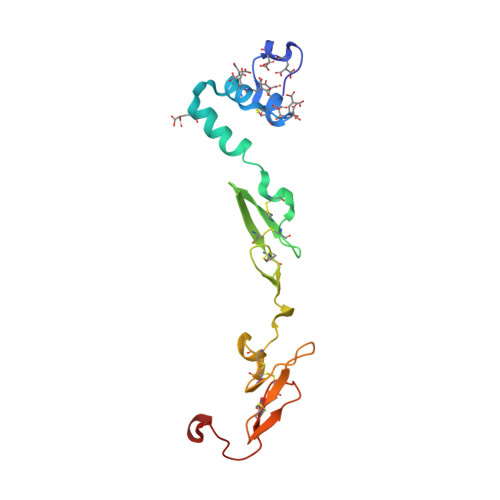
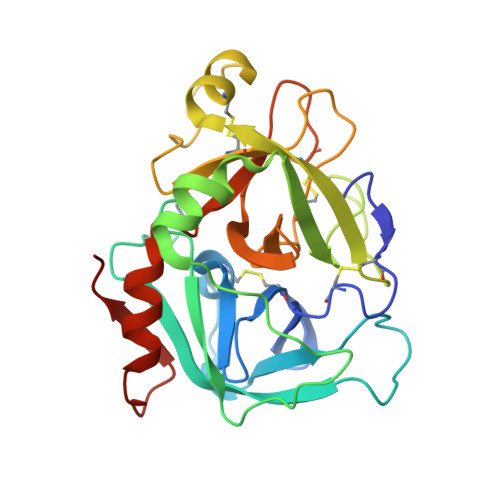
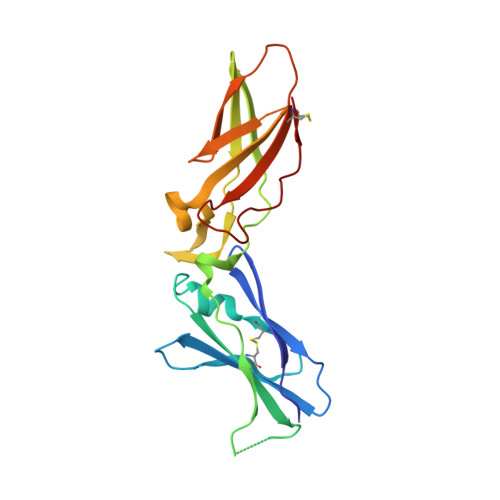
Mg2+ Is Required for Optimal Folding of the Gamma-Carboxyglutamic Acid (Gla) Domains of Vitamin K-Dependent Clotting Factors At Physiological Ca2+
Vadivel, K., Agah, S., Messer, A., Cascio, D., Bajaj, S.M., Krishnaswamy, S., Esmon, C.T., Padmanabhan, K., Bajaj, S.P.To be published.