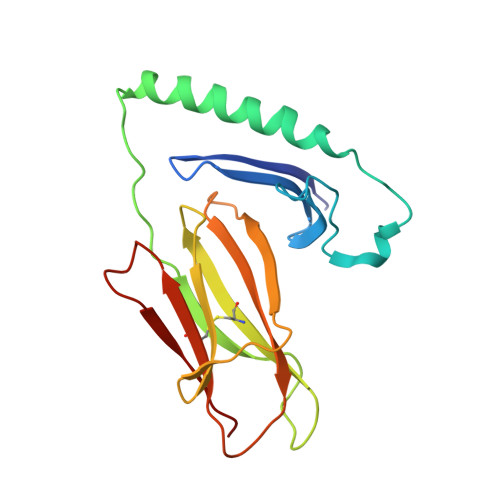
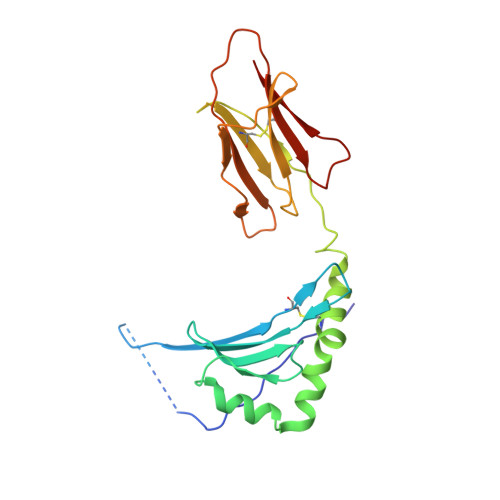
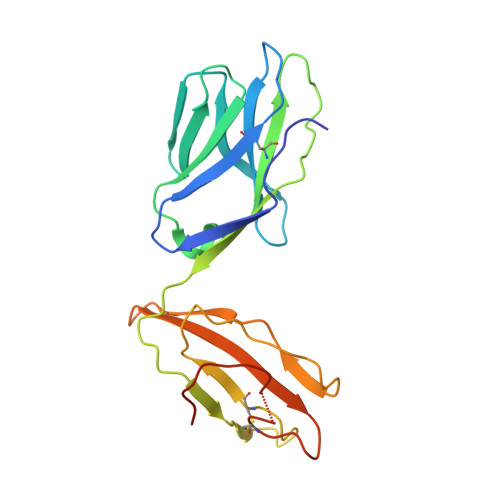
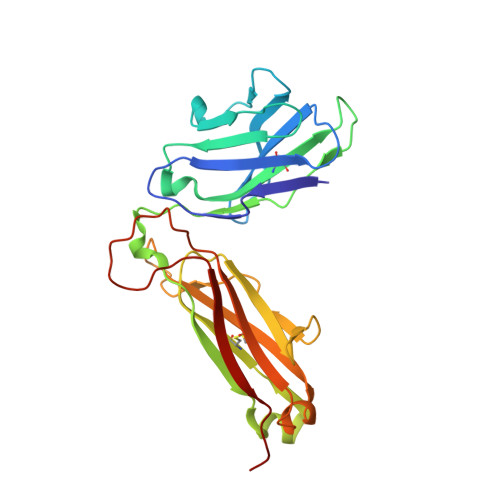
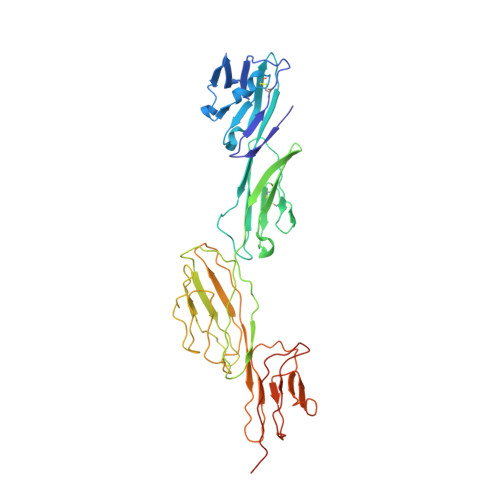
Crystal structure of a complete ternary complex of T-cell receptor, peptide-MHC, and CD4.
Yin, Y., Wang, X.X., Mariuzza, R.A.(2012) Proc Natl Acad Sci U S A 109: 5405-5410
- PubMed: 22431638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1118801109
- Primary Citation Related Structures:
3T0E - PubMed Abstract:
Adaptive immunity depends on specific recognition by a T-cell receptor (TCR) of an antigenic peptide bound to a major histocompatibility complex (pMHC) molecule on an antigen-presenting cell (APC). In addition, T-cell activation generally requires binding of this same pMHC to a CD4 or CD8 coreceptor. Here, we report the structure of a complete TCR-pMHC-CD4 ternary complex involving a human autoimmune TCR, a myelin-derived self-peptide bound to HLA-DR4, and CD4. The complex resembles a pointed arch in which TCR and CD4 are each tilted ∼65° relative to the T-cell membrane. By precluding direct contacts between TCR and CD4, the structure explains how TCR and CD4 on the T cell can simultaneously, yet independently, engage the same pMHC on the APC. The structure, in conjunction with previous mutagenesis data, places TCR-associated CD3εγ and CD3εδ subunits, which transmit activation signals to the T cell, inside the TCR-pMHC-CD4 arch, facing CD4. By establishing anchor points for TCR and CD4 on the T-cell membrane, the complex provides a basis for understanding how the CD4 coreceptor focuses TCR on MHC to guide TCR docking on pMHC during thymic T-cell selection.
- W. M. Keck Laboratory for Structural Biology, Institute for Bioscience and Biotechnology Research, University of Maryland, Rockville, MD 20850, USA.
Organizational Affiliation: