Determination of engineered chloride-binding site structures in fluorescent proteins reveals principles of halide recognition
Wang, W., Grimley, J.S., Augustine, G.J., Beese, L.S., Hellinga, H.W.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Green fluorescent protein | 258 | Aequorea victoria | Mutation(s): 6 Gene Names: GFP | 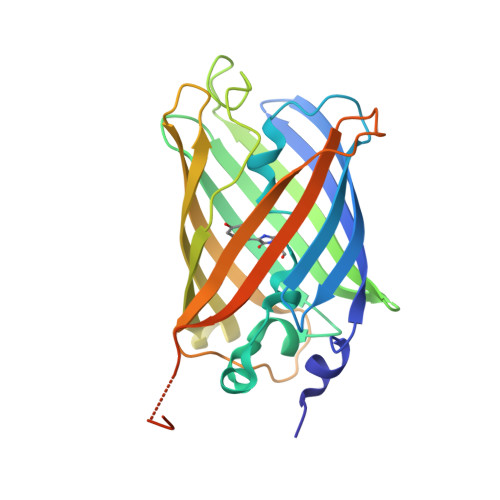 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P42212 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| IOD Download:Ideal Coordinates CCD File | B [auth A], C [auth A], D [auth A], E [auth A], F [auth A] | IODIDE ION I XMBWDFGMSWQBCA-UHFFFAOYSA-M |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CR2 Query on CR2 | A | L-PEPTIDE LINKING | C13 H13 N3 O4 |  | GLY, TYR, GLY |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.297 | α = 90 |
| b = 62.741 | β = 90 |
| c = 69.783 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| SERGUI | data collection |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |