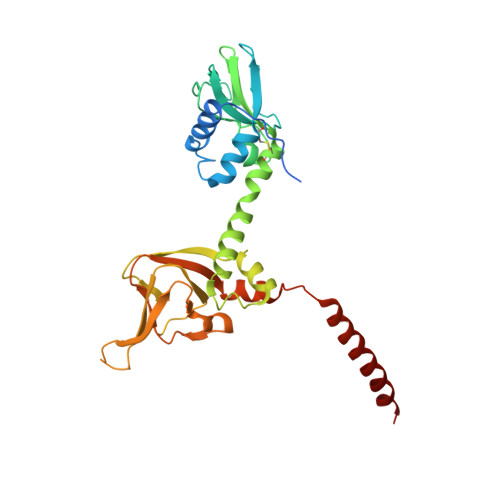
Crystal Structure of cGMP-Dependent Protein Kinase Reveals Novel Site of Interchain Communication.
Osborne, B.W., Wu, J., McFarland, C.J., Nickl, C.K., Sankaran, B., Casteel, D.E., Woods, V.L., Kornev, A.P., Taylor, S.S., Dostmann, W.R.(2011) Structure 19: 1317-1327
- PubMed: 21893290
- DOI: https://doi.org/10.1016/j.str.2011.06.012
- Primary Citation of Related Structures:
3SHR - PubMed Abstract:
The cGMP-dependent protein kinase (PKG) serves as an integral component of second messenger signaling in a number of biological contexts including cell differentiation, memory, and vasodilation. PKG is homodimeric and large conformational changes accompany cGMP binding. However, the structure of PKG and the molecular mechanisms associated with protomer communication following cGMP-induced activation remain unknown. Here, we report the 2.5 Å crystal structure of a regulatory domain construct (aa 78-355) containing both cGMP binding sites of PKG Iα. A distinct and segregated architecture with an extended central helix separates the two cGMP binding domains. Additionally, a previously uncharacterized helical domain (switch helix) promotes the formation of a hydrophobic interface between protomers. Mutational disruption of this interaction in full-length PKG implicates the switch helix as a critical site of dimer communication in PKG biology. These results offer new structural insight into the mechanism of allosteric PKG activation.
- Department of Pharmacology, College of Medicine, University of Vermont, Burlington, VT 05405, USA.
Organizational Affiliation: