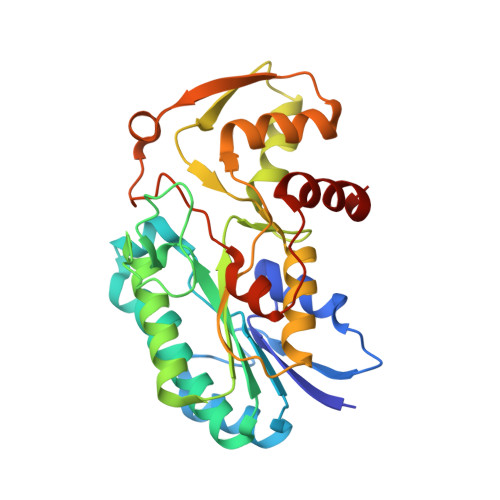
Structure of the Bacillus anthracis dTDP-L-rhamnose-biosynthetic enzyme dTDP-4-dehydrorhamnose reductase (RfbD).
Law, A., Stergioulis, A., Halavaty, A.S., Minasov, G., Anderson, W.F., Kuhn, M.L.(2017) Acta Crystallogr F Struct Biol Commun 73: 644-650
- PubMed: 29199984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17015746
- Primary Citation Related Structures:
3SC6 - PubMed Abstract:
Bacillus anthracis is the causative agent of the deadly disease Anthrax. Its use in bioterrorism and its ability to re-emerge have brought renewed interest in this organism. B. anthracis is a Gram-positive bacterium that adds L-rhamnose to its cell-wall polysaccharides using the activated donor dTDP-β-L-rhamnose. The enzymes involved in the biosynthesis of the activated donor are absent in humans, which make them ideal targets for therapeutic development to combat pathogens. Here, the 2.65 Å resolution crystal structure of the fourth enzyme in the dTDP-β-L-rhamnose-biosynthetic pathway from B. anthracis, dTDP-4-dehydro-β-L-rhamnose reductase (RfbD), is presented in complex with NADP + . This enzyme catalyzes the reduction of dTDP-4-dehydro-β-L-rhamnose to dTDP-β-L-rhamnose. Although the protein was co-crystallized in the presence of Mg 2+ , the protein lacks the conserved residues that coordinate Mg 2+ .
- Department of Chemistry and Biochemistry, San Francisco State University, USA.
Organizational Affiliation: