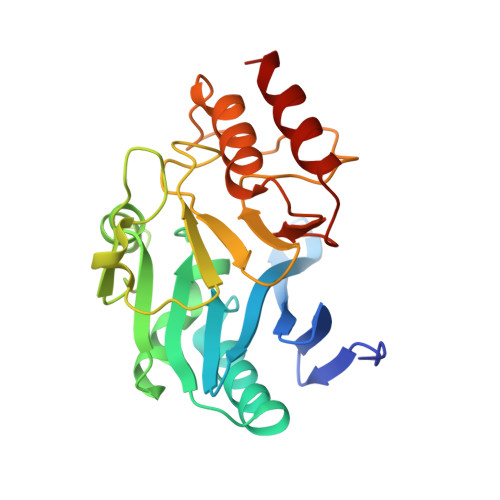
Structure of Apo- and Monometalated Forms of NDM-1 A Highly Potent Carbapenem-Hydrolyzing Metallo-beta-Lactamase
Kim, Y., Tesar, C., Mire, J., Jedrzejczak, R., Binkowski, A., Babnigg, G., Sacchettini, J., Joachimiak, A.(2011) PLoS One 6: e24621-e24621
- PubMed: 21931780 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0024621
- Primary Citation Related Structures:
3RKJ, 3RKK, 3SBL, 3SFP - PubMed Abstract:
The New Delhi Metallo-β-lactamase (NDM-1) gene makes multiple pathogenic microorganisms resistant to all known β-lactam antibiotics. The rapid emergence of NDM-1 has been linked to mobile plasmids that move between different strains resulting in world-wide dissemination. Biochemical studies revealed that NDM-1 is capable of efficiently hydrolyzing a wide range of β-lactams, including many carbapenems considered as "last resort" antibiotics. The crystal structures of metal-free apo- and monozinc forms of NDM-1 presented here revealed an enlarged and flexible active site of class B1 metallo-β-lactamase. This site is capable of accommodating many β-lactam substrates by having many of the catalytic residues on flexible loops, which explains the observed extended spectrum activity of this zinc dependent β-lactamase. Indeed, five loops contribute "keg" residues in the active site including side chains involved in metal binding. Loop 1 in particular, shows conformational flexibility, apparently related to the acceptance and positioning of substrates for cleavage by a zinc-activated water molecule.
- Midwest Center for Structural Genomics and Structural Biology Center, Biosciences, Argonne National Laboratory, Argonne, Illinois, United States of America.
Organizational Affiliation: