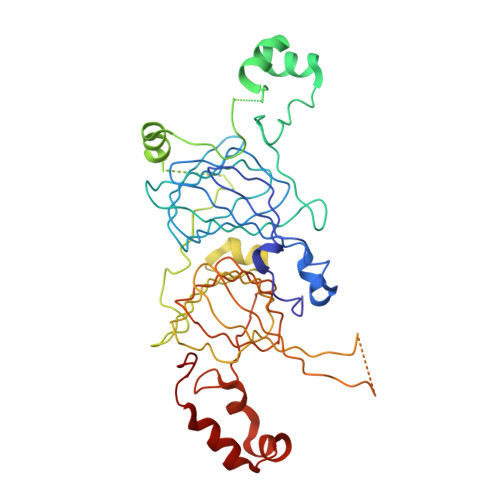
Structural and Immunologic Characterization of Ara h 1, a Major Peanut Allergen.
Chruszcz, M., Maleki, S.J., Majorek, K.A., Demas, M., Bublin, M., Solberg, R., Hurlburt, B.K., Ruan, S., Mattisohn, C.P., Breiteneder, H., Minor, W.(2011) J Biological Chem 286: 39318-39327
- PubMed: 21917921 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.270132
- Primary Citation Related Structures:
3S7E, 3S7I - PubMed Abstract:
Allergic reactions to peanuts and tree nuts are major causes of anaphylaxis in the United States. We compare different properties of natural and recombinant versions of Ara h 1, a major peanut allergen, through structural, immunologic, and bioinformatics analyses. Small angle x-ray scattering studies show that natural Ara h 1 forms higher molecular weight aggregates in solution. In contrast, the full-length recombinant protein is partially unfolded and exists as a monomer. The crystal structure of the Ara h 1 core (residues 170-586) shows that the central part of the allergen has a bicupin fold, which is in agreement with our bioinformatics analysis. In its crystalline state, the core region of Ara h 1 forms trimeric assemblies, while in solution the protein exists as higher molecular weight assemblies. This finding reveals that the residues forming the core region of the protein are sufficient for formation of Ara h 1 trimers and higher order oligomers. Natural and recombinant variants of proteins tested in in vitro gastric and duodenal digestion assays show that the natural protein is the most stable form, followed by the recombinant Ara h 1 core fragment and the full-length recombinant protein. Additionally, IgE binding studies reveal that the natural and recombinant allergens have different patterns of interaction with IgE antibodies. The molecular basis of cross-reactivity between vicilin allergens is also elucidated.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, Virginia 22908, USA. maks@iwonka.med.virginia.edu
Organizational Affiliation: