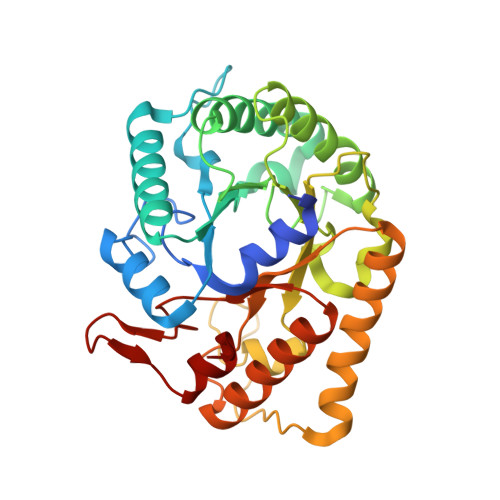
Crystal structure of hyperthermophilic Endo-beta-1,4-glucanase: Implications for catalytic mechanism and thermostability.
Zheng, B., Yang, W., Zhao, X., Wang, Y., Lou, Z., Rao, Z., Feng, Y.(2012) J Biological Chem 287: 8336-8346
- PubMed: 22128157 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.266346
- Primary Citation Related Structures:
3RJX, 3RJY - PubMed Abstract:
Endo-β-1,4-glucanase from thermophilic Fervidobacterium nodosum Rt17-B1 (FnCel5A), a new member of glycosyl hydrolase family 5, is highly thermostable and exhibits the highest activity on carboxymethylcellulose among the reported homologues. To understand the structural basis for the thermostability and catalytic mechanism, we report here the crystal structures of FnCel5A and the complex with glucose at atomic resolution. FnCel5A exhibited a (β/α)(8)-barrel structure typical of clan GH-A of the glycoside hydrolase families with a large and deep catalytic pocket located in the C-terminal end of the β-strands that may permit substrate access. A comparison of the structure of FnCel5A with related structures from thermopile Clostridium thermocellum, mesophile Clostridium cellulolyticum, and psychrophile Pseudoalteromonas haloplanktis showed significant differences in intramolecular interactions (salt bridges and hydrogen bonds) that may account for the difference in their thermostabilities. The substrate complex structure in combination with a mutagenesis analysis of the catalytic residues implicates a distinctive catalytic module Glu(167)-His(226)-Glu(283), which suggests that the histidine may function as an intermediate for the electron transfer network between the typical Glu-Glu catalytic module. Further investigation suggested that the aromatic residues Trp(61), Trp(204), Phe(231), and Trp(240) as well as polar residues Asn(51), His(127), Tyr(228), and His(235) in the active site not only participated in substrate binding but also provided a unique microenvironment suitable for catalysis. These results provide substantial insight into the unique characteristics of FnCel5A for catalysis and adaptation to extreme temperature.
- Key Laboratory for Molecular Enzymology and Engineering of Ministry of Education, Jilin University, Changchun 130023, China.
Organizational Affiliation: