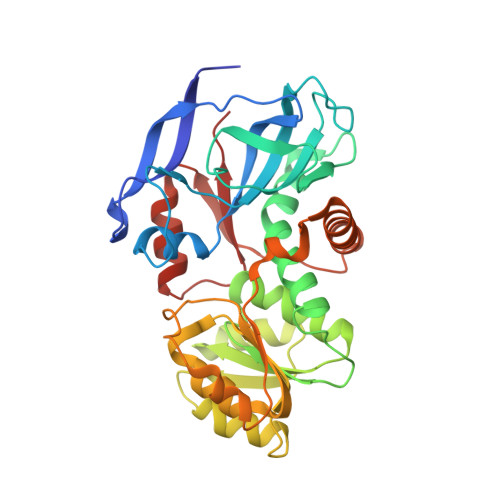
Structural insights into the cofactor-assisted substrate recognition of yeast quinone oxidoreductase Zta1
Guo, P.C., Ma, X.X., Bao, Z.Z., Ma, J.D., Chen, Y., Zhou, C.Z.(2011) J Struct Biol 176: 112-118
- PubMed: 21820057 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2011.07.010
- Primary Citation Related Structures:
3QWA, 3QWB - PubMed Abstract:
Quinone oxidoreductase (QOR EC1.6.5.5) catalyzes the reduction of quinone to hydroxyquinone using NADPH as a cofactor. Here we present the crystal structure of the ζ-crystallin-like QOR Zta1 from Saccharomycescerevisiae in apo-form at 2.00 Å and complexed with NADPH at 1.59 Å resolution. Zta1 forms a homodimer, with each subunit containing a catalytic and a cofactor-binding domain. Upon NADPH binding to the interdomain cleft, the two domains shift towards each other, producing a better fit for NADPH, and tightening substrate binding. Computational simulation combined with site-directed mutagenesis and enzymatic activity analysis defined a potential quinone-binding site that determines the stringent substrate specificity. Moreover, multiple-sequence alignment and kinetics assays implied that a single-residue change from Arg in lower organisms to Gly in vertebrates possibly resulted in elevation of enzymatic activity of ζ-crystallin-like QORs throughout evolution.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei Anhui 230027, People's Republic of China.
Organizational Affiliation: