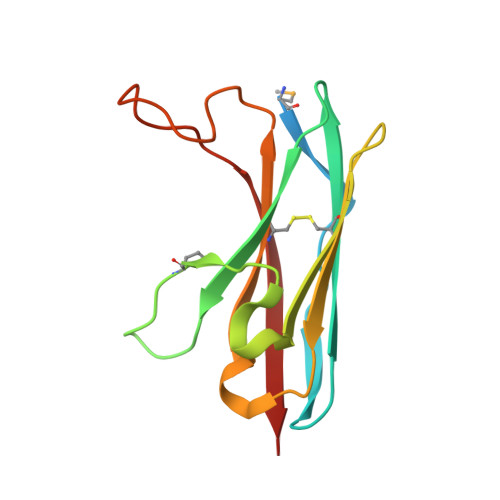
The Retinal Specific CD147 Ig0 Domain: From Molecular Structure to Biological Activity.
Redzic, J.S., Armstrong, G.S., Isern, N.G., Jones, D.N., Kieft, J.S., Eisenmesser, E.Z.(2011) J Mol Biology 411: 68-82
- PubMed: 21620857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.04.060
- Primary Citation Related Structures:
3QQN, 3QR2 - PubMed Abstract:
CD147 is a type I transmembrane protein that is involved in inflammatory diseases, cancer progression, and multiple human pathogens utilize CD147 for efficient infection. CD147 expression is so high in several cancers that it is now used as a prognostic marker. The two primary isoforms of CD147 that are related to cancer progression have been identified, differing in their number of immunoglobulin (Ig)-like domains. These include CD147 Ig1-Ig2, which is ubiquitously expressed in most tissues, and CD147 Ig0-Ig1-Ig2, which is retinal specific and implicated in retinoblastoma. However, little is known in regard to the retinal specific CD147 Ig0 domain despite its potential role in retinoblastoma. We present the first crystal structure of the human CD147 Ig0 domain and show that the CD147 Ig0 domain is a crystallographic dimer with an I-type domain structure, which maintained in solution. Furthermore, we have utilized our structural data together with mutagenesis to probe the biological activity of CD147-containing proteins, both with and without the CD147 Ig0 domain, within several model cell lines. Our findings reveal that the CD147 Ig0 domain is a potent stimulator of interleukin-6 and suggest that the CD147 Ig0 domain has its own receptor distinct from that of the other CD147 Ig-like domains, CD147 Ig1-Ig2. Finally, we show that the CD147 Ig0 dimer is the functional unit required for activity and can be disrupted by a single point mutation.
- Department of Biochemistry and Molecular Genetics, University of Colorado Denver, School of Medicine, Aurora, CO 80045, USA.
Organizational Affiliation: