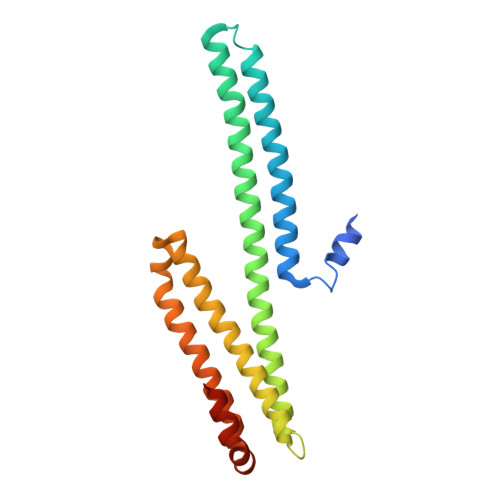
Crystal structure of amyloid precursor-like protein 1 and heparin complex suggests a dual role of heparin in E2 dimerization.
Xue, Y., Lee, S., Ha, Y.(2011) Proc Natl Acad Sci U S A 108: 16229-16234
- PubMed: 21930949 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1103407108
- Primary Citation Related Structures:
3QMK - PubMed Abstract:
Mutations in amyloid precursor protein (APP) are associated with familial Alzheimer's disease. Recent development suggests that homo- and heterodimerization of APP and APP-like proteins (APLPs), which are enhanced by heparan sulfate binding, may play a role in signal transduction and cell adhesion. Despite efforts to model heparin binding based on known apo crystal structures, the mechanism of heparin-induced APP/APLP dimerization has not been established experimentally. Here we report the crystal structure of a complex between heparin and the E2 domain of APLP1, which harbors the conserved high affinity heparin binding site of the full-length molecule. Within the asymmetric E2:heparin complex, the polysaccharide is snugly bound inside a narrow groove between the two helical subdomains of one protein protomer. The nonreducing end of the sugar is positioned near the protein's 2-fold axis, making contacts with basic residues from the second protomer. The inability of the E2 dimer to accommodate two heparin molecules near its symmetry axis explains the observed 21 binding stoichiometry, which is confirmed by isothermal titration calorimetric experiment carried out in solution. We also show that, at high concentrations, heparin can destabilize E2 dimer, probably by forcing into the unoccupied binding site observed in the 21 complex. The binding model suggested by the crystal structure may facilitate the design of heparin mimetics that are capable of modulating APP dimerization in cells.
- Department of Pharmacology, Yale School of Medicine, New Haven, CT 06520, USA.
Organizational Affiliation: