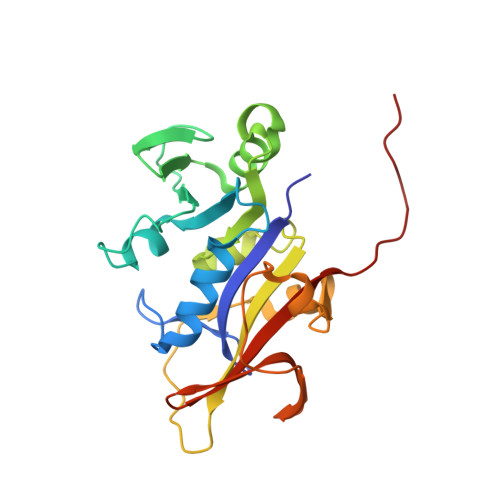
Crystal Structures of Candida albicans Dihydrofolate Reductase Bound to Propargyl-Linked Antifolates Reveal the Flexibility of Active Site Loop Residues Critical for Ligand Potency and Selectivity.
Paulsen, J.L., Bendel, S.D., Anderson, A.C.(2011) Chem Biol Drug Des 78: 505-512
- PubMed: 21726415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1747-0285.2011.01169.x
- Primary Citation Related Structures:
3QLR, 3QLS, 3QLW, 3QLX, 3QLY, 3QLZ - PubMed Abstract:
Candida albicans and Candida glabrata cause fungal bloodstream infections that are associated with significant mortality. As part of an effort to develop potent and selective antifolates that target dihydrofolate reductase (DHFR) from Candida species, we report three ternary crystal structures of C. albicans DHFR (CaDHFR) bound to novel propargyl-linked analogs. Consistent with earlier modeling results, these structures show that hydrophobic pockets in the binding site may be exploited to increase ligand potency. The crystal structures also confirm that loop residues Thr 58- Phe 66, which flank the active site and influence ligand potency and selectivity, adopt multiple conformations. To aid the development of a dual Candida spp. inhibitor, three new crystal structures of C. glabrata DHFR (CgDHFR) bound to similar ligands as those bound in the ternary structures of CaDHFR are also reported here. Loop residues 58-66 in CgDHFR and human DHFR are 1 and 3 Å closer to the folate binding site, respectively, than loop residues in CaDHFR, suggesting that a properly size ligand could be a potent and selective dual inhibitor of CaDHFR and CgDHFR.
- Department of Pharmaceutical Sciences, University of Connecticut, Storrs, CT 06269, USA.
Organizational Affiliation: