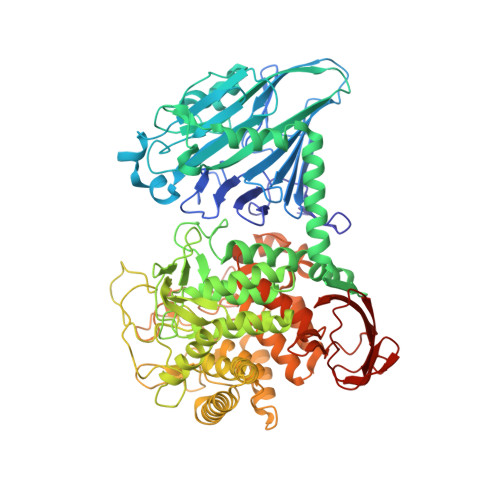
Structure of cellobiose phosphorylase from Clostridium thermocellum in complex with phosphate.
Bianchetti, C.M., Elsen, N.L., Fox, B.G., Phillips, G.N.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1345-1349
- PubMed: 22102229 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111032660
- Primary Citation Related Structures:
3QDE - PubMed Abstract:
Clostridium thermocellum is a cellulosome-producing bacterium that is able to efficiently degrade and utilize cellulose as a sole carbon source. Cellobiose phosphorylase (CBP) plays a critical role in cellulose degradation by catalyzing the reversible phosphate-dependent hydrolysis of cellobiose, the major product of cellulose degradation, into α-D-glucose 1-phosphate and D-glucose. CBP from C. thermocellum is a modular enzyme composed of four domains [N-terminal domain, helical linker, (α/α)(6)-barrel domain and C-terminal domain] and is a member of glycoside hydrolase family 94. The 2.4 Å resolution X-ray crystal structure of C. thermocellum CBP reveals the residues involved in coordinating the catalytic phosphate as well as the residues that are likely to be involved in substrate binding and discrimination.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, WI 53706, USA.
Organizational Affiliation: