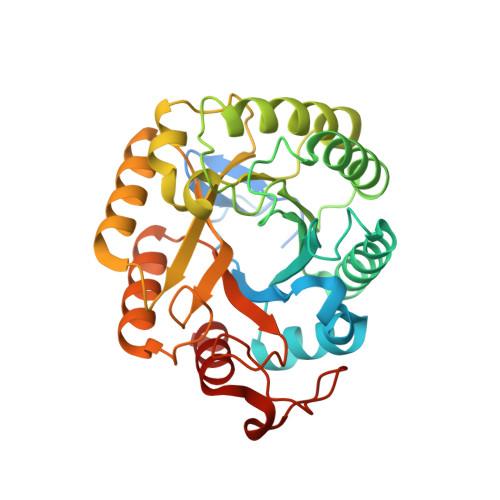
Dissecting structure-function-stability relationships of a thermostable GH5-CBM3 cellulase from Bacillus subtilis 168.
Santos, C.R., Paiva, J.H., Sforca, M.L., Neves, J.L., Navarro, R.Z., Cota, J., Akao, P.K., Hoffmam, Z.B., Meza, A.N., Smetana, J.H., Nogueira, M.L., Polikarpov, I., Xavier-Neto, J., Squina, F.M., Ward, R.J., Ruller, R., Zeri, A.C., Murakami, M.T.(2012) Biochem J 441: 95-104
- PubMed: 21880019 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20110869
- Primary Citation Related Structures:
2L8A, 3PZT, 3PZU, 3PZV - PubMed Abstract:
Cellulases participate in a number of biological events, such as plant cell wall remodelling, nematode parasitism and microbial carbon uptake. Their ability to depolymerize crystalline cellulose is of great biotechnological interest for environmentally compatible production of fuels from lignocellulosic biomass. However, industrial use of cellulases is somewhat limited by both their low catalytic efficiency and stability. In the present study, we conducted a detailed functional and structural characterization of the thermostable BsCel5A (Bacillus subtilis cellulase 5A), which consists of a GH5 (glycoside hydrolase 5) catalytic domain fused to a CBM3 (family 3 carbohydrate-binding module). NMR structural analysis revealed that the Bacillus CBM3 represents a new subfamily, which lacks the classical calcium-binding motif, and variations in NMR frequencies in the presence of cellopentaose showed the importance of polar residues in the carbohydrate interaction. Together with the catalytic domain, the CBM3 forms a large planar surface for cellulose recognition, which conducts the substrate in a proper conformation to the active site and increases enzymatic efficiency. Notably, the manganese ion was demonstrated to have a hyper-stabilizing effect on BsCel5A, and by using deletion constructs and X-ray crystallography we determined that this effect maps to a negatively charged motif located at the opposite face of the catalytic site.
- Laboratório Nacional de Biociências, Centro Nacional de Pesquisa em Energia e Materiais, Campinas SP, Brazil.
Organizational Affiliation: