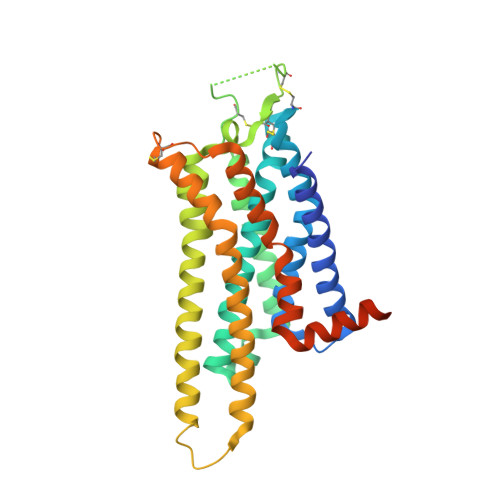
Structure of the adenosine A(2A) receptor in complex with ZM241385 and the xanthines XAC and caffeine
Dore, A.S., Robertson, N., Errey, J.C., Ng, I., Hollenstein, K., Tehan, B., Hurrell, E., Bennett, K., Congreve, M., Magnani, F., Tate, C.G., Weir, M., Marshall, F.H.(2011) Structure 19: 1283-1293
- PubMed: 21885291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.06.014
- Primary Citation Related Structures:
3PWH, 3REY, 3RFM - PubMed Abstract:
Methylxanthines, including caffeine and theophylline, are among the most widely consumed stimulant drugs in the world. These effects are mediated primarily via blockade of adenosine receptors. Xanthine analogs with improved properties have been developed as potential treatments for diseases such as Parkinson's disease. Here we report the structures of a thermostabilized adenosine A(2A) receptor in complex with the xanthines xanthine amine congener and caffeine, as well as the A(2A) selective inverse agonist ZM241385. The receptor is crystallized in the inactive state conformation as defined by the presence of a salt bridge known as the ionic lock. The complete third intracellular loop, responsible for G protein coupling, is visible consisting of extended helices 5 and 6. The structures provide new insight into the features that define the ligand binding pocket of the adenosine receptor for ligands of diverse chemotypes as well as the cytoplasmic regions that interact with signal transduction proteins.
- Heptares Therapeutics Ltd, BioPark, Welwyn Garden City, Herts, AL7 3AX, UK.
Organizational Affiliation: