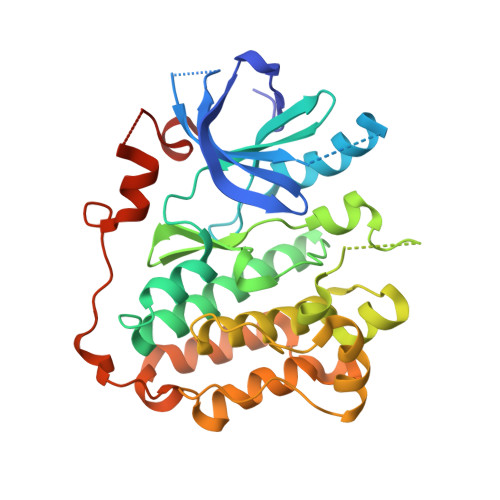
Structural Analysis of the Mechanism of Inhibition and Allosteric Activation of the Kinase Domain of HER2 Protein.
Aertgeerts, K., Skene, R., Yano, J., Sang, B.C., Zou, H., Snell, G., Jennings, A., Iwamoto, K., Habuka, N., Hirokawa, A., Ishikawa, T., Tanaka, T., Miki, H., Ohta, Y., Sogabe, S.(2011) J Biological Chem 286: 18756-18765
- PubMed: 21454582 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.206193
- Primary Citation Related Structures:
3POZ, 3PP0 - PubMed Abstract:
Aberrant signaling of ErbB family members human epidermal growth factor 2 (HER2) and epidermal growth factor receptor (EGFR) is implicated in many human cancers, and HER2 expression is predictive of human disease recurrence and prognosis. Small molecule kinase inhibitors of EGFR and of both HER2 and EGFR have received approval for the treatment of cancer. We present the first high resolution crystal structure of the kinase domain of HER2 in complex with a selective inhibitor to understand protein activation, inhibition, and function at the molecular level. HER2 kinase domain crystallizes as a dimer and suggests evidence for an allosteric mechanism of activation comparable with previously reported activation mechanisms for EGFR and HER4. A unique Gly-rich region in HER2 following the α-helix C is responsible for increased conformational flexibility within the active site and could explain the low intrinsic catalytic activity previously reported for HER2. In addition, we solved the crystal structure of the kinase domain of EGFR in complex with a HER2/EGFR dual inhibitor (TAK-285). Comparison with previously reported inactive and active EGFR kinase domain structures gave insight into the mechanism of HER2 and EGFR inhibition and may help guide the design and development of new cancer drugs with improved potency and selectivity.
- Takeda San Diego Inc, 10410 Science Center Drive, San Diego, California 92121, USA. kaertgeerts@takedasd.com
Organizational Affiliation: