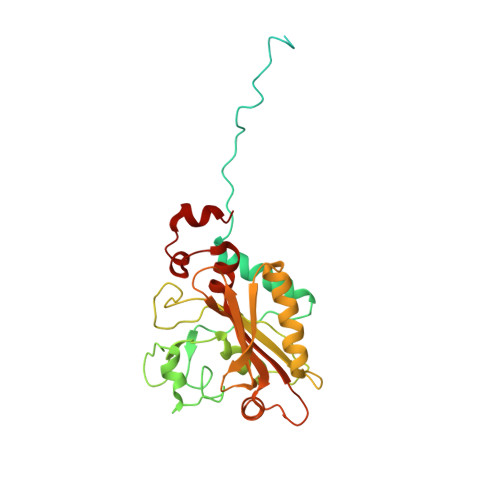
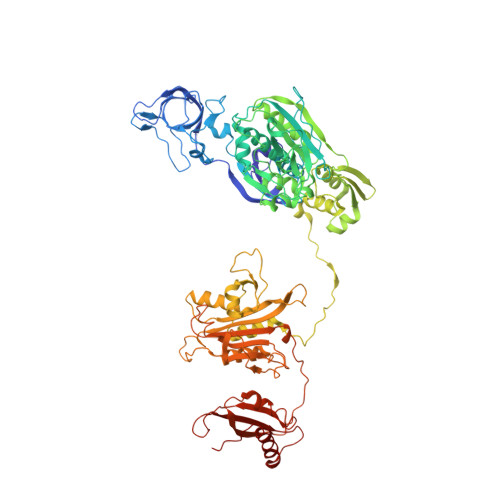
Idiosyncrasy and identity in the prokaryotic phe-system: crystal structure of E. coli phenylalanyl-tRNA synthetase complexed with phenylalanine and AMP.
Mermershtain, I., Finarov, I., Klipcan, L., Kessler, N., Rozenberg, H., Safro, M.G.(2011) Protein Sci 20: 160-167
- PubMed: 21082706
- DOI: https://doi.org/10.1002/pro.549
- Primary Citation of Related Structures:
3PCO - PubMed Abstract:
The crystal structure of Phenylalanyl-tRNA synthetase from E. coli (EcPheRS), a class II aminoacyl-tRNA synthetase, complexed with phenylalanine and AMP was determined at 3.05 Å resolution. EcPheRS is a (αβ)₂ heterotetramer: the αβ heterodimer of EcPheRS consists of 11 structural domains. Three of them: the N-terminus, A1 and A2 belong to the α-subunit and B1-B8 domains to the β subunit. The structure of EcPheRS revealed that architecture of four helix-bundle interface, characteristic of class IIc heterotetrameric aaRSs, is changed: each of the two long helices belonging to CLM transformed into the coil-short helix structural fragments. The N-terminal domain of the α-subunit in EcPheRS forms compact triple helix domain. This observation is contradictory to the structure of the apo form of TtPheRS, where N-terminal domain was not detected in the electron density map. Comparison of EcPheRS structure with TtPheRS has uncovered significant rearrangements of the structural domains involved in tRNA(Phe) binding/translocation. As it follows from modeling experiments, to achieve a tighter fit with anticodon loop of tRNA, a shift of ∼5 Å is required for C-terminal domain B8, and of ∼6 to 7 Å for the whole N terminus. EcPheRSs have emerged as an important target for the incorporation of novel amino acids into genetic code. Further progress in design of novel compounds is anticipated based on the structural data of EcPheRS.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: