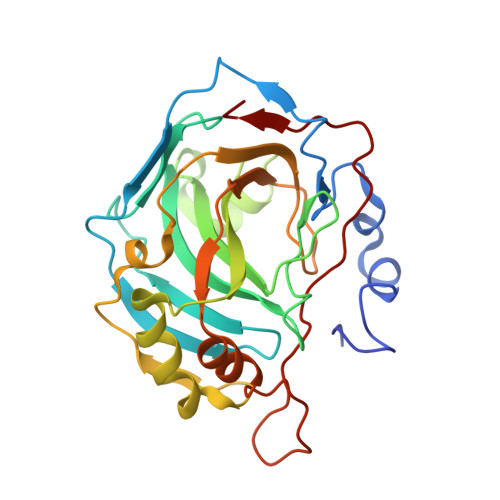
Protein crystal structures with ferrocene and ruthenocene-based enzyme inhibitors.
Salmon, A.J., Williams, M.L., Hofmann, A., Poulsen, S.A.(2012) Chem Commun (Camb) 48: 2328-2330
- PubMed: 22258283 Search on PubMed
- DOI: https://doi.org/10.1039/c2cc15625c
- Primary Citation Related Structures:
3P3H, 3P3J, 3P44, 3P55 - PubMed Abstract:
We have determined the protein X-ray crystal structures of four organometallic inhibitors in complex with their target enzyme carbonic anhydrase II. The barrel-shaped hydrophobic ferrocene and ruthenocene moieties have provided a structure-based avenue to better occupy the hydrophobic binding patch within the enzyme active site.
- Eskitis Institute for Cell and Molecular Therapies, Griffith University, Nathan, Queensland 4111, Australia.
Organizational Affiliation: