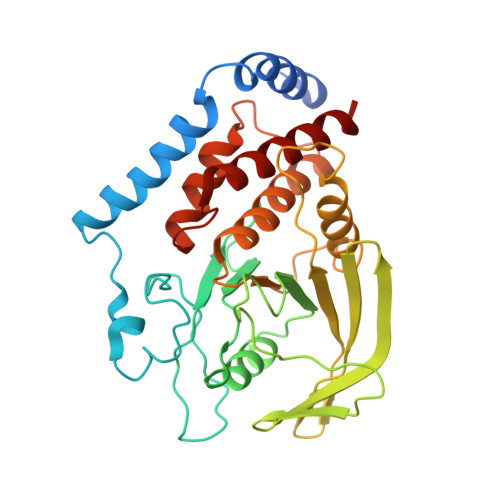
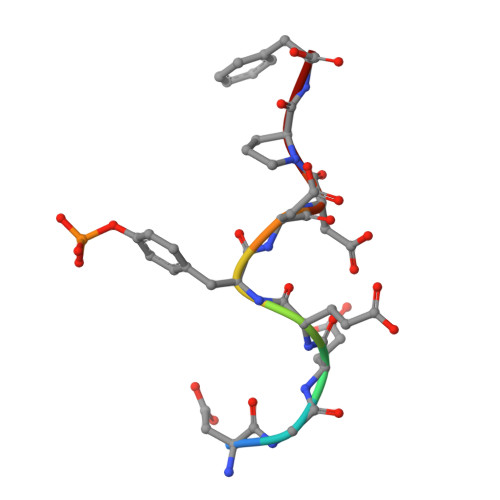
Substrate Specificity of Lymphoid-specific Tyrosine Phosphatase (Lyp) and Identification of Src Kinase-associated Protein of 55 kDa Homolog (SKAP-HOM) as a Lyp Substrate.
Yu, X., Chen, M., Zhang, S., Yu, Z.H., Sun, J.P., Wang, L., Liu, S., Imasaki, T., Takagi, Y., Zhang, Z.Y.(2011) J Biological Chem 286: 30526-30534
- PubMed: 21719704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.254722
- Primary Citation Related Structures:
3OMH - PubMed Abstract:
A missense single-nucleotide polymorphism in the gene encoding the lymphoid-specific tyrosine phosphatase (Lyp) has been identified as a causal factor in a wide spectrum of autoimmune diseases. Interestingly, the autoimmune-predisposing variant of Lyp appears to represent a gain-of-function mutation, implicating Lyp as an attractive target for the development of effective strategies for the treatment of many autoimmune disorders. Unfortunately, the precise biological functions of Lyp in signaling cascades and cellular physiology are poorly understood. Identification and characterization of Lyp substrates will help define the chain of molecular events coupling Lyp dysfunction to diseases. In the current study, we identified consensus sequence motifs for Lyp substrate recognition using an "inverse alanine scanning" combinatorial library approach. The intrinsic sequence specificity data led to the discovery and characterization of SKAP-HOM, a cytosolic adaptor protein required for proper activation of the immune system, as a bona fide Lyp substrate. To determine the molecular basis for Lyp substrate recognition, we solved crystal structures of Lyp in complex with the consensus peptide as well as the phosphopeptide derived from SKAP-HOM. Together with the biochemical data, the structures define the molecular determinants for Lyp substrate specificity and provide a solid foundation upon which novel therapeutics targeting Lyp can be developed for multiple autoimmune diseases.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana 46202.
Organizational Affiliation: