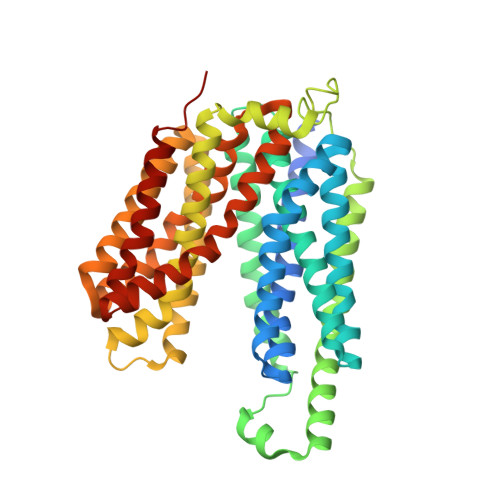
Structure of a fucose transporter in an outward-open conformation
Dang, S.Y., Sun, L.F., Huang, Y., Lu, F., Liu, Y., Gong, H., Wang, J., Yan, N.(2010) Nature 467: 734-738
- PubMed: 20877283 Search on PubMed
- DOI: https://doi.org/10.1038/nature09406
- Primary Citation Related Structures:
3O7P, 3O7Q - PubMed Abstract:
The major facilitator superfamily (MFS) transporters are an ancient and widespread family of secondary active transporters. In Escherichia coli, the uptake of l-fucose, a source of carbon for microorganisms, is mediated by an MFS proton symporter, FucP. Despite intensive study of the MFS transporters, atomic structure information is only available on three proteins and the outward-open conformation has yet to be captured. Here we report the crystal structure of FucP at 3.1 Å resolution, which shows that it contains an outward-open, amphipathic cavity. The similarly folded amino and carboxyl domains of FucP have contrasting surface features along the transport path, with negative electrostatic potential on the N domain and hydrophobic surface on the C domain. FucP only contains two acidic residues along the transport path, Asp 46 and Glu 135, which can undergo cycles of protonation and deprotonation. Their essential role in active transport is supported by both in vivo and in vitro experiments. Structure-based biochemical analyses provide insights into energy coupling, substrate recognition and the transport mechanism of FucP.
- State Key Laboratory of Bio-membrane and Membrane Biotechnology, Center for Structural Biology, School of Life Sciences and School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: