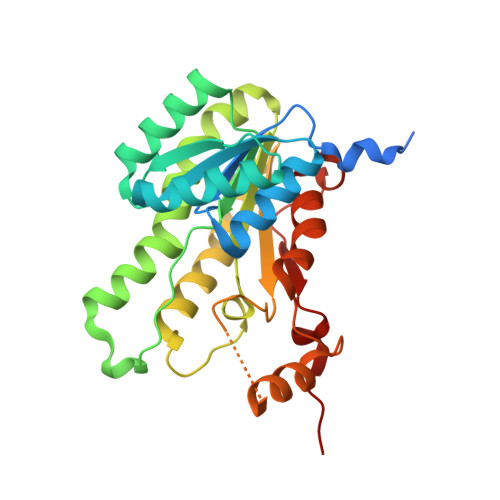
Structural Insight Into the Catalytic Mechanism of Gluconate 5-Dehydrogenase from Streptococcus Suis: Crystal Structures of the Substrate-Free and Quaternary Complex Enzymes.
Zhang, Q., Peng, H., Gao, F., Liu, Y., Cheng, H., Thompson, J., Gao, G.F.(2009) Protein Sci 18: 294
- PubMed: 19177572 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.32
- Primary Citation Related Structures:
3CXR, 3O03 - PubMed Abstract:
Gluconate 5-dehydrogenase (Ga5DH) is an NADP(H)-dependent enzyme that catalyzes a reversible oxidoreduction reaction between D-gluconate and 5-keto-D-gluconate, thereby regulating the flux of this important carbon and energy source in bacteria. Despite the considerable amount of physiological and biochemical knowledge of Ga5DH, there is little physical or structural information available for this enzyme. To this end, we herein report the crystal structures of Ga5DH from pathogenic Streptococcus suis serotype 2 in both substrate-free and liganded (NADP(+)/D-gluconate/metal ion) quaternary complex forms at 2.0 A resolution. Structural analysis reveals that Ga5DH adopts a protein fold similar to that found in members of the short chain dehydrogenase/reductase (SDR) family, while the enzyme itself represents a previously uncharacterized member of this family. In solution, Ga5DH exists as a tetramer that comprised four identical approximately 29 kDa subunits. The catalytic site of Ga5DH shows considerable architectural similarity to that found in other enzymes of the SDR family, but the S. suis protein contains an additional residue (Arg104) that plays an important role in the binding and orientation of substrate. The quaternary complex structure provides the first clear crystallographic evidence for the role of a catalytically important serine residue and also reveals an amino acid tetrad RSYK that differs from the SYK triad found in the majority of SDR enzymes. Detailed analysis of the crystal structures reveals important contributions of Ca(2+) ions to active site formation and of specific residues at the C-termini of subunits to tetramer assembly. Because Ga5DH is a potential target for therapy, our findings provide insight not only of catalytic mechanism, but also suggest a target of structure-based drug design.
- Institute of Microbiology, Chinese Academy of Sciences, Beijing, People's Republic Of China.
Organizational Affiliation: