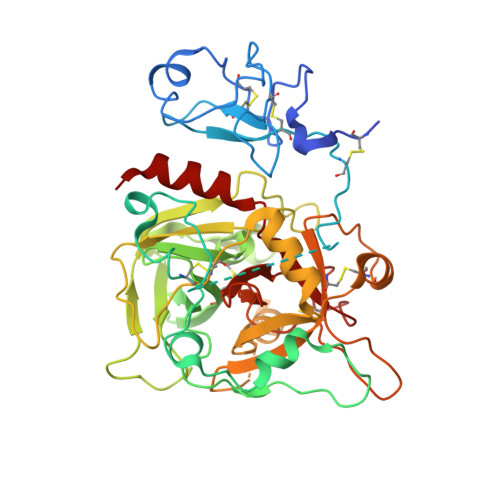
Crystal structure of prethrombin-1.
Chen, Z., Pelc, L.A., Di Cera, E.(2010) Proc Natl Acad Sci U S A 107: 19278-19283
- PubMed: 20974933 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1010262107
- Primary Citation Related Structures:
3NXP - PubMed Abstract:
Prothrombin is the zymogen precursor of the clotting enzyme thrombin, which is generated by two sequential cleavages at R271 and R320 by the prothrombinase complex. The structure of prothrombin is currently unknown. Prethrombin-1 differs from prothrombin for the absence of 155 residues in the N-terminal domain and is composed of a single polypeptide chain containing fragment 2 (residues 156-271), A chain (residues 272-320), and B chain (residues 321-579). The X-ray crystal structure of prethrombin-1 solved at 2.2-Å resolution shows an overall conformation significantly different (rmsd = 3.6 Å) from that of its active form meizothrombin desF1 carrying a cleavage at R320. Fragment 2 is rotated around the y axis by 29° and makes only few contacts with the B chain. In the B chain, the oxyanion hole is disrupted due to absence of the I16-D194 ion pair and the Na(+) binding site and adjacent primary specificity pocket are highly perturbed. A remarkable feature of the structure is that the autolysis loop assumes a helical conformation enabling W148 and W215, located 17 Å apart in meizothrombin desF1, to come within 3.3 Å of each other and completely occlude access to the active site. These findings suggest that the zymogen form of thrombin possesses conformational plasticity comparable to that of the mature enzyme and have significant implications for the mechanism of prothrombin activation and the zymogen → protease conversion in trypsin-like proteases.
- Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, Saint Louis, MO 63104, USA.
Organizational Affiliation: