Crystal Structure of Human Peripheral Myelin Protein 2
Ugochukwu, E., Pilka, E., Phillips, C., Yue, W.W., Krojer, T., von Delft, F., Bountra, C., Arrowsmith, C.H., Weigelt, J., Edwards, A., Kavanagh, K.L., Oppermann, U.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| PMP2 protein | 153 | Homo sapiens | Mutation(s): 0 Gene Names: hCG_21300, PMP2 | 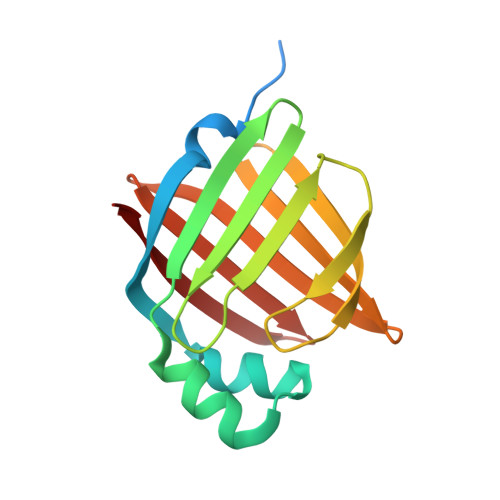 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P02689 GTEx: ENSG00000147588 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P02689 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PLM Download:Ideal Coordinates CCD File | B [auth A] | PALMITIC ACID C16 H32 O2 IPCSVZSSVZVIGE-UHFFFAOYSA-N |  | ||
| EPE Download:Ideal Coordinates CCD File | H [auth A] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| CIT Download:Ideal Coordinates CCD File | G [auth A] | CITRIC ACID C6 H8 O7 KRKNYBCHXYNGOX-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | C [auth A], D [auth A], E [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | F [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | I [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 67.19 | α = 90 |
| b = 67.19 | β = 90 |
| c = 100.07 | γ = 90 |
| Software Name | Purpose |
|---|---|
| MAR345 | data collection |
| PHASER | phasing |
| REFMAC | refinement |
| MOSFLM | data reduction |
| SCALA | data scaling |