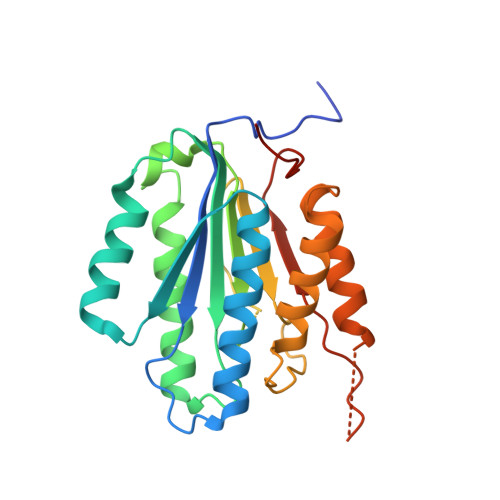
Substrate-Induced Conformational Changes Occur in All Cleaved Forms of Caspase-6.
Vaidya, S., Velazquez-Delgado, E.M., Abbruzzese, G., Hardy, J.A.(2011) J Mol Biology 406: 75-91
- PubMed: 21111746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.11.031
- Primary Citation Related Structures:
3NKF - PubMed Abstract:
Caspase-6 is an apoptotic cysteine protease that also governs disease progression in Huntington's and Alzheimer's diseases. Caspase-6 is of great interest as a target for treatment of these neurodegenerative diseases; however, the molecular basis of caspase-6 function and regulation remains poorly understood. In the recently reported structure of caspase-6, the 60's and 130's helices at the base of the substrate-binding groove extend upward, in a conformation entirely different from that of any other caspase. Presently, the central question about caspase-6 structure and function is whether the extended conformation is the catalytically competent conformation or whether the extended helices must undergo a large conformational rearrangement in order to bind substrate. We have generated a series of caspase-6 cleavage variants, including a novel constitutively two-chain form, and determined crystal structures of caspase-6 with and without the intersubunit linker. This series allows evaluation of the role of the prodomain and intersubunit linker on caspase-6 structure and function before and after substrate binding. Caspase-6 is inherently more stable than closely related caspases. Cleaved caspase-6 with both the prodomain and the linker present is the most stable, indicating that these two regions act in concert to increase stability, but maintain the extended conformation in the unliganded state. Moreover, these data suggest that caspase-6 undergoes a significant conformational change upon substrate binding, adopting a structure that is more like canonical caspases.
- Department of Chemistry, University of Massachusetts Amherst, 104 Lederle Graduate Research Tower,710 North Pleasant Street, Amherst, MA 01003, USA.
Organizational Affiliation: