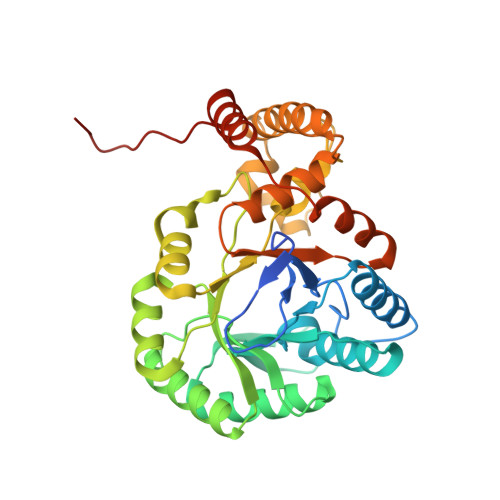
The Three-Dimensional Structure of AKR11B4, a Glycerol Dehydrogenase from Gluconobacter oxydans, Reveals a Tryptophan Residue as an Accelerator of Reaction Turnover.
Richter, N., Breicha, K., Hummel, W., Niefind, K.(2010) J Mol Biology 404: 353-362
- PubMed: 20887732
- DOI: https://doi.org/10.1016/j.jmb.2010.09.049
- Primary Citation Related Structures:
3N2T - PubMed Abstract:
The NADP-dependent glycerol dehydrogenase (EC 1.1.1.72) from Gluconobacter oxydans is a member of family 11 of the aldo-keto reductase (AKR) enzyme superfamily; according to the systematic nomenclature within the AKR superfamily, the term AKR11B4 has been assigned to the enzyme. AKR11B4 is a biotechnologically attractive enzyme because of its broad substrate spectrum, combined with its distinctive regioselectivity and stereoselectivity. These features can be partially rationalized based on a 2-Å crystal structure of apo-AKR11B4, which we describe and interpret here against the functional complex structures of other members of family 11 of the AKR superfamily. The structure of AKR11B4 shows the AKR-typical (β/α)(8) TIM-barrel fold, with three loops and the C-terminal tail determining the particular enzymatic properties. In comparison to AKR11B1 (its closest AKR relative), AKR11B4 has a relatively broad binding cleft for the cosubstrate NADP/NADPH. In the crystalline environment, it is completely blocked by the C-terminal segment of a neighboring protomer. The structure reveals a conspicuous tryptophan residue (Trp23) that has to adopt an unconventional and strained side-chain conformation to permit cosubstrate binding. We predict and confirm by site-directed mutagenesis that Trp23 is an accelerator of (co)substrate turnover. Furthermore, we show that, simultaneously, this tryptophan residue is a critical determinant for substrate binding by the enzyme, while enantioselectivity is probably governed by a methionine residue within the C-terminal tail. We present structural reasons for these notions based on ternary complex models of AKR11B4, NADP, and either octanal, d-glyceraldehyde, or l-glyceraldehyde.
- Evocatal GmbH, Merowingerplatz 1A, D-40225 Düsseldorf, Germany.
Organizational Affiliation: