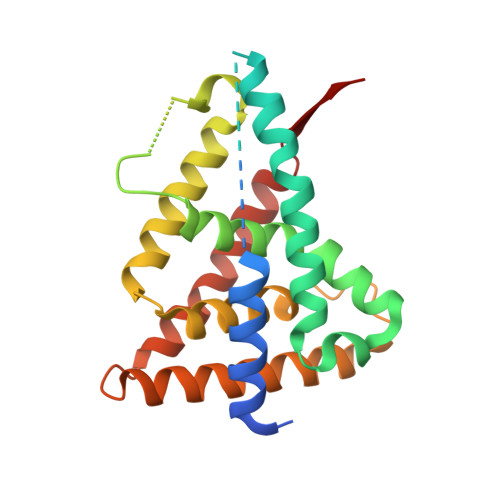
Structure of Rev-erbalpha bound to N-CoR reveals a unique mechanism of nuclear receptor-co-repressor interaction.
Phelan, C.A., Gampe, R.T., Lambert, M.H., Parks, D.J., Montana, V., Bynum, J., Broderick, T.M., Hu, X., Williams, S.P., Nolte, R.T., Lazar, M.A.(2010) Nat Struct Mol Biol 17: 808-814
- PubMed: 20581824 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1860
- Primary Citation Related Structures:
3N00 - PubMed Abstract:
Repression of gene transcription by the nuclear receptor Rev-erbalpha plays an integral role in the core molecular circadian clock. We report the crystal structure of a nuclear receptor-co-repressor (N-CoR) interaction domain 1 (ID1) peptide bound to truncated human Rev-erbalpha ligand-binding domain (LBD). The ID1 peptide forms an unprecedented antiparallel beta-sheet with Rev-erbalpha, as well as an alpha-helix similar to that seen in nuclear receptor ID2 crystal structures but out of register by four residues. Comparison with the structure of Rev-erbbeta bound to heme indicates that ID1 peptide and heme induce substantially different conformational changes in the LBD. Although heme is involved in Rev-erb repression, the structure suggests that Rev-erbalpha could also mediate repression via ID1 binding in the absence of heme. The previously uncharacterized secondary structure induced by ID1 peptide binding advances our understanding of nuclear receptor-co-repressor interactions.
- Division of Endocrinology, Diabetes and Metabolism, Departments of Medicine and Genetics, The Institute for Diabetes, Obesity and Metabolism, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania, USA.
Organizational Affiliation: