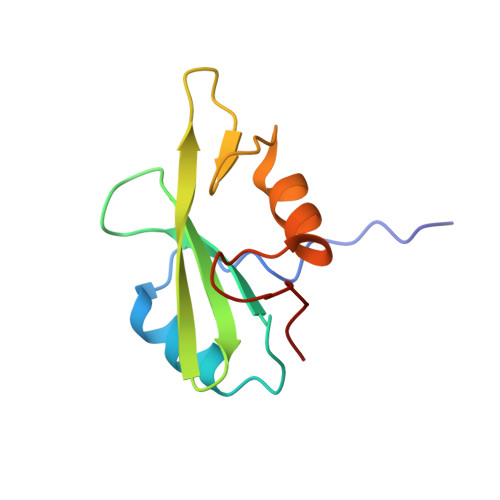
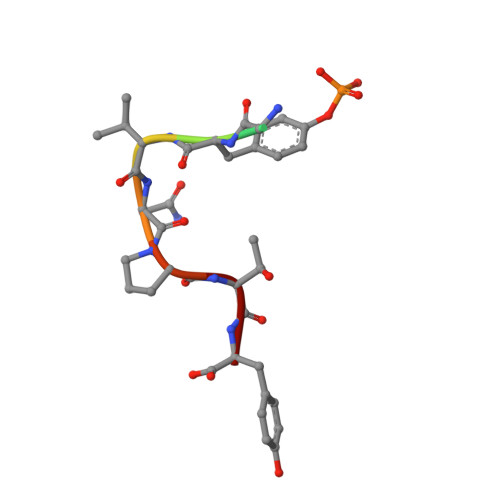
Functional Implications of the Conformational Switch in AICD Peptide upon Binding to Grb2-SH2 Domain.
Das, S., Raychaudhuri, M., Sen, U., Mukhopadhyay, D.(2011) J Mol Biology 414: 217-230
- PubMed: 22001015 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.09.046
- Primary Citation Related Structures:
3MXC, 3MXY - PubMed Abstract:
It has been hypothesized previously that synergistic effect of both amyloid precursor protein intracellular C-terminal domain (AICD) and Aβ aggregation could contribute to Alzheimer's disease pathogenesis. Structural studies of AICD have found no stable globular fold over a broad range of pH. Present work is based on the premises that a conformational switch involving the flipping of C-terminal helix of AICD would be essential for effective binding with the Src homology 2 (SH2) domain of growth factor receptor binding protein-2 (Grb2) and subsequent initiation of Grb2-mediated endo-lysosomal pathway. High-resolution crystal structures of Grb2-SH2 domain bound to AICD peptides reveal a unique mode of binding where the peptides assume a noncanonical conformation that is unlike other structures of AICD peptides bound to protein-tyrosine-binding domains or that of its free state; rather, a flipping of the C-terminal helix of AICD is evident. The involvement of different AICD residues in Grb2-SH2 interaction is further elucidated through fluorescence-based assays. Our results reveal the significance of a specific interaction of the two molecules to optimize the rapid transport of AICD inside endosomal vesicles presumably to reduce the cytotoxic load.
- Structural Genomics Division, Saha Institute of Nuclear Physics, 1/AF Bidhan Nagar, WB, Kolkata 700 064, India.
Organizational Affiliation: