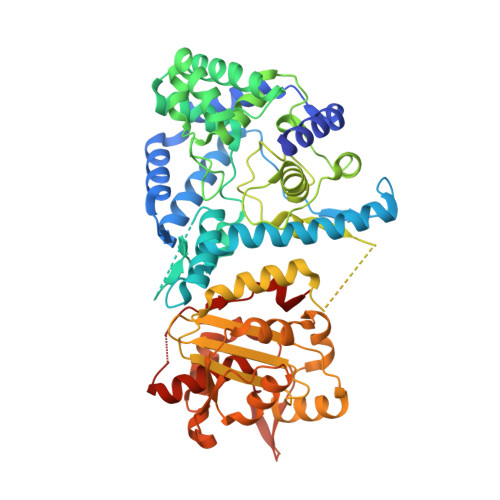
Cap binding and immune evasion revealed by Lassa nucleoprotein structure.
Qi, X., Lan, S., Wang, W., Schelde, L.M., Dong, H., Wallat, G.D., Ly, H., Liang, Y., Dong, C.(2010) Nature 468: 779-783
- PubMed: 21085117 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09605
- Primary Citation Related Structures:
3MWP, 3MWT, 3MX2, 3MX5 - PubMed Abstract:
Lassa virus, the causative agent of Lassa fever, causes thousands of deaths annually and is a biological threat agent, for which there is no vaccine and limited therapy. The nucleoprotein (NP) of Lassa virus has essential roles in viral RNA synthesis and immune suppression, the molecular mechanisms of which are poorly understood. Here we report the crystal structure of Lassa virus NP at 1.80 Å resolution, which reveals amino (N)- and carboxy (C)-terminal domains with structures unlike any of the reported viral NPs. The N domain folds into a novel structure with a deep cavity for binding the m7GpppN cap structure that is required for viral RNA transcription, whereas the C domain contains 3'-5' exoribonuclease activity involved in suppressing interferon induction. To our knowledge this is the first X-ray crystal structure solved for an arenaviral NP, which reveals its unexpected functions and indicates unique mechanisms in cap binding and immune evasion. These findings provide great potential for vaccine and drug development.
- Biomedical Sciences Research Complex, School of Chemistry, University of St Andrews, North Haugh, St Andrews, Fife KY16 9ST, UK.
Organizational Affiliation: