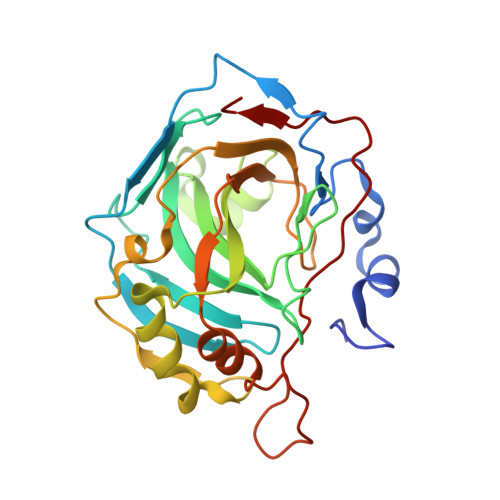
Carbonic anhydrase inhibitors. The X-ray crystal structure of human isoform II in adduct with an adamantyl analogue of acetazolamide resides in a less utilized binding pocket than most hydrophobic inhibitors.
Avvaru, B.S., Wagner, J.M., Maresca, A., Scozzafava, A., Robbins, A.H., Supuran, C.T., McKenna, R.(2010) Bioorg Med Chem Lett 20: 4376-4381
- PubMed: 20605094 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.06.082
- Primary Citation Related Structures:
3MHC - PubMed Abstract:
We investigated the inhibitory activity of several 1,3,4-thiadiazole-sulfonamides against all catalytically active CA (EC 4.2.1.1), CA I-XV. The tail derivatizing the 5-position in the 1,3,4-thiadiazole-2-sulfonamide scaffold was observed to be critical as an inhibitory determinant of these compounds. The high resolution X-ray crystal structure of hCA II in complex with 5-(1-adamantylcarboxamido)-1,3,4-thiadiazole-2-sulfonamide, showed the adamantyl moiety of the inhibitor residing in a less utilized binding pocket than that of most hydrophobic inhibitors, lined by the amino acid residues Ile91, Val121 and Phe131. This binding site may explain the diverse inhibition profiles of 5-carboxamide- and sufonamide-derivatized 1,3,4-thiadiazole-2-sulfonamides and offers a hot spot for designing isoform selective inhibitors, considering that residues 91 and 131 are highly variable among the 13 catalytically active isoforms.
- Department of Biochemistry and Molecular Biology, College of Medicine, University of Florida, Box Gainesville, FL 32610, USA.
Organizational Affiliation: