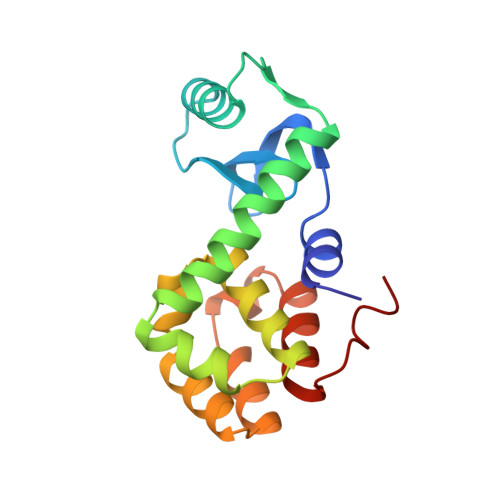
Structural studies of mutants of T4 lysozyme that alter hydrophobic stabilization.
Matsumura, M., Wozniak, J.A., Sun, D.P., Matthews, B.W.(1989) J Biological Chem 264: 16059-16066
- PubMed: 2674124 Search on PubMed
- Primary Citation Related Structures:
3LZM - PubMed Abstract:
Multiple replacements at amino acid position 3 of bacteriophage T4 lysozyme have shown that the conformational stability of the protein is directly governed by the hydrophobicity of the residue substituted (Matsumura, M., Becktel, W. J., and Matthews, B. W. (1988) Nature 334, 406-410). Of the 13 mutant lysozymes made by site-directed mutagenesis, two variants, one with valine (I3V) and the other with tyrosine (I3Y), were crystallized and their structures solved. In this report we describe the crystal structures of these variants at 1.7 A resolution. While the structure of the I3V mutant is essentially the same as that of wild-type lysozyme, the I3Y mutant has substantial changes in its structure. The most significant of these are that the side chain of the tyrosine is not accommodated within the interior of the protein and the amino-terminal polypeptide (residues 1-9) moves 0.6-1.1 A relative to the wild-type structure. Using coordinates based on the wild-type and available mutant structures, solvent accessible surface area of residue 3 as well as the adjacent 9 residues in the folded form were calculated. The free energy of stabilization based on the transfer of these residues from a fully extended form to the interior to the folded protein was found to correlate well with the protein stability determined by thermodynamic analysis. The enhanced thermostability of the variant Ile-3----Leu, relative to wild-type lysozyme, can also be rationalized by surface-area calculations based on a model-built structure. Noncrystallization of most lysozyme variants at position 3 appears to be due to disruption of intermolecular contacts in the crystal. The Ile-3----Val variant is closely isomorphous with wild-type and maintains the same crystal contacts. In the Ile-3----Tyr variant, however, a new set of contacts is made in which direct protein-protein hydrogen bonds are replaced by protein-water-protein hydrogen bonds as well as a novel hydrogen bond involving the phenolic hydroxyl of the substituted tyrosine.
- Institute of Molecular Biology, University of Oregon, Eugene 97403.
Organizational Affiliation: