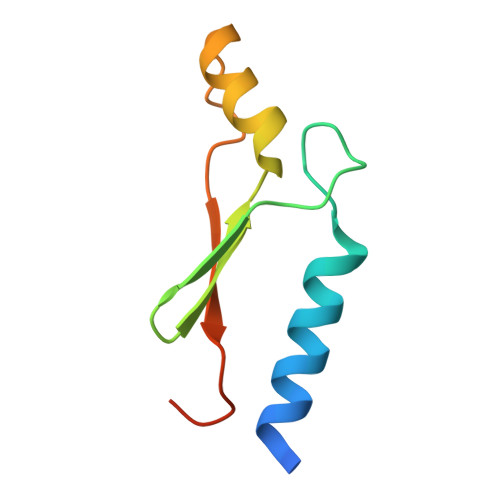
Three-dimensional structures reveal multiple ADP/ATP binding modes for a synthetic class of artificial proteins.
Simmons, C.R., Magee, C.L., Smith, D.A., Lauman, L., Chaput, J.C., Allen, J.P.(2010) Biochemistry 49: 8689-8699
- PubMed: 20822107 Search on PubMed
- DOI: https://doi.org/10.1021/bi100398p
- Primary Citation Related Structures:
3LT8, 3LT9, 3LTA, 3LTB, 3LTC, 3LTD - PubMed Abstract:
The creation of synthetic enzymes with predefined functions represents a major challenge in future synthetic biology applications. Here, we describe six structures of de novo proteins that have been determined using protein crystallography to address how simple enzymes perform catalysis. Three structures are of a protein, DX, selected for its stability and ability to tightly bind ATP. Despite the addition of ATP to the crystallization conditions, the presence of a bound but distorted ATP was found only under excess ATP conditions, with ADP being present under equimolar conditions or when crystallized for a prolonged period of time. A bound ADP cofactor was evident when Asp was substituted for Val at residue 65, but ATP in a linear configuration is present when Phe was substituted for Tyr at residue 43. These new structures complement previously determined structures of DX and the protein with the Phe 43 to Tyr substitution [Simmons, C. R., et al. (2009) ACS Chem. Biol. 4, 649-658] and together demonstrate the multiple ADP/ATP binding modes from which a model emerges in which the DX protein binds ATP in a configuration that represents a transitional state for the catalysis of ATP to ADP through a slow, metal-free reaction capable of multiple turnovers. This unusual observation suggests that design-free methods can be used to generate novel protein scaffolds that are tailor-made for catalysis.
- Center for Evolutionary Medicine and Informatics, The Biodesign Institute, Arizona State University, Tempe, Arizona 85287, USA.
Organizational Affiliation: