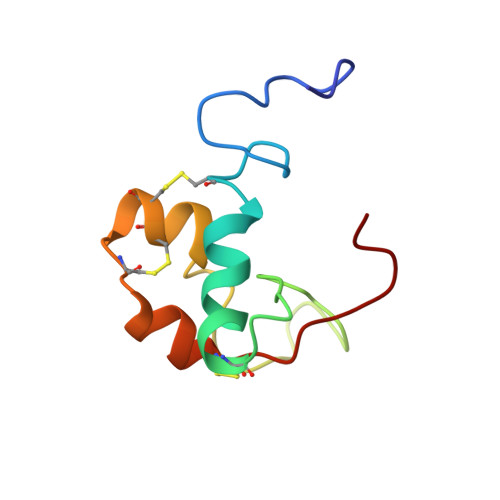
Solution structure and backbone dynamics of long-[Arg(3)]insulin-like growth factor-I
Laajoki, L.G., Francis, G.L., Wallace, J.C., Carver, J.A., Keniry, M.A.(2000) J Biological Chem 275: 10009-10015
- PubMed: 10744677 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.275.14.10009
- Primary Citation Related Structures:
3LRI - PubMed Abstract:
Long-[Arg(3)]insulin-like growth factor-I (IGF-I) is a potent analog of insulin-like growth factor-I that has been modified by a Glu(3) --> Arg mutation and a 13-amino acid extension appended to the N terminus. We have determined the solution structure of (15)N-labeled Long-[Arg(3)]-IGF-I using high resolution NMR and restrained molecular dynamics techniques to a precision of 0.82 +/- 0.28 A root mean square deviation for the backbone heavy atoms in the three alpha-helices and 3.5 +/- 0.9 A root mean square deviation for all backbone heavy atoms excluding the 8 N-terminal residues and the 8 C-terminal eight residues. Overall, the structure of the IGF-I domain is consistent with earlier studies of IGF-I with some minor changes remote from the N terminus. The major variations in the structure, compared with IGF-I, occur at the N terminus with a substantial reorientation of the N-terminal three residues of the IGF-I domain. These results are interpreted in terms of the lower binding affinity for insulin-like growth factor-binding proteins. The backbone dynamics of Long-[Arg(3)]IGF-I were investigated using (15)N nuclear spin relaxation and the heteronuclear nuclear Overhauser enhancement (NOE). There is a considerable degree of flexibility in Long-[Arg(3)]IGF-I, even in the alpha-helices, as indicated by an average ((1)H)(15)N NOE of 0.55 for the regions. The largest heteronuclear NOEs are observed in the helical regions, lower heteronuclear NOEs are observed in the C-domain loop separating helix 1 from helix 2, and negative heteronuclear NOEs are observed in the N-terminal extension and at the C terminus. Despite these data indicating conformational flexibility for the N-terminal extension, slow amide proton exchange was observed for some residues in this region, suggesting some transitory structure does exist, possibly a molten helix. A certain degree of flexibility may be necessary in all insulin-like growth factors to enable association with various receptors and binding proteins.
- Research School of Chemistry, The Australian National University, Canberra, Australian Capital Territory 2601, South Australia 5000.
Organizational Affiliation: