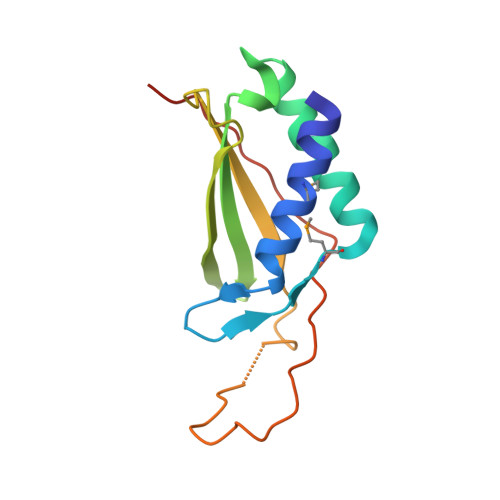
Structural analysis and functional implications of the negative mTORC1 regulator REDD1.
Vega-Rubin-de-Celis, S., Abdallah, Z., Kinch, L., Grishin, N.V., Brugarolas, J., Zhang, X.(2010) Biochemistry 49: 2491-2501
- PubMed: 20166753 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi902135e
- Primary Citation Related Structures:
3LQ9 - PubMed Abstract:
REDD1 is a conserved stress-response protein that regulates mTORC1, a critical regulator of cell growth and proliferation that is implicated in cancer. REDD1 is induced by hypoxia, and REDD1 overexpression is sufficient to inhibit mTORC1. mTORC1 is regulated by the small GTPase Rheb, which in turn is regulated by the GTPase-activating protein complex, TSC1/TSC2. REDD1 induced-mTORC1 inhibition requires the TSC1/TSC2 complex, and REDD1 has been proposed to act by directly binding to and sequestering 14-3-3 proteins away from TSC2 leading to TSC2-dependent inhibition of mTORC1. Structure/function analyses have led us to identify two segments in REDD1 that are essential for function, which act in an interdependent manner. We have determined a crystal structure of REDD1 at 2.0 A resolution, which shows that these two segments fold together to form an intact domain with a novel fold. This domain is characterized by an alpha/beta sandwich consisting of two antiparallel alpha-helices and a mixed beta-sheet encompassing an uncommon psi-loop motif. Structure-based docking and functional analyses suggest that REDD1 does not directly bind to 14-3-3 proteins. Sequence conservation mapping to the surface of the structure and mutagenesis studies demarcated a hotspot likely to interact with effector proteins that is essential for REDD1-mediated mTORC1 inhibition.
- Department of Developmental Biology, University of Texas Southwestern Medical Center, Dallas, Texas 75390, USA.
Organizational Affiliation: