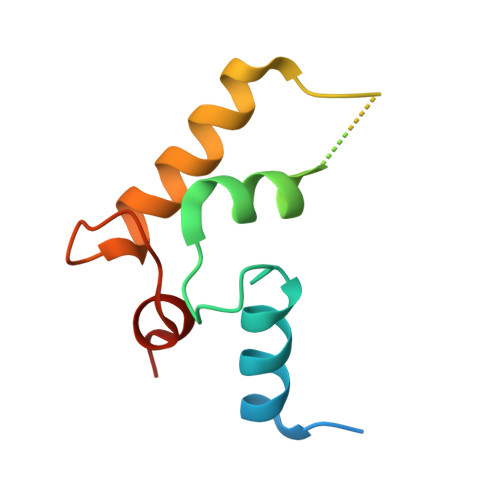
Crystal structure of the LMAN1-CRD/MCFD2 transport receptor complex provides insight into combined deficiency of factor V and factor VIII.
Wigren, E., Bourhis, J.M., Kursula, I., Guy, J.E., Lindqvist, Y.(2010) FEBS Lett 584: 878-882
- PubMed: 20138881 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2010.02.009
- Primary Citation Related Structures:
3LCP - PubMed Abstract:
LMAN1 is a glycoprotein receptor, mediating transfer from the ER to the ER-Golgi intermediate compartment. Together with the co-receptor MCFD2, it transports coagulation factors V and VIII. Mutations in LMAN1 and MCFD2 can cause combined deficiency of factors V and VIII (F5F8D). We present the crystal structure of the LMAN1/MCFD2 complex and relate it to patient mutations. Circular dichroism data show that the majority of the substitution mutations give rise to a disordered or severely destabilized MCFD2 protein. The few stable mutation variants are found in the binding surface of the complex leading to impaired LMAN1 binding and F5F8D.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, 17177 Stockholm, Sweden.
Organizational Affiliation: