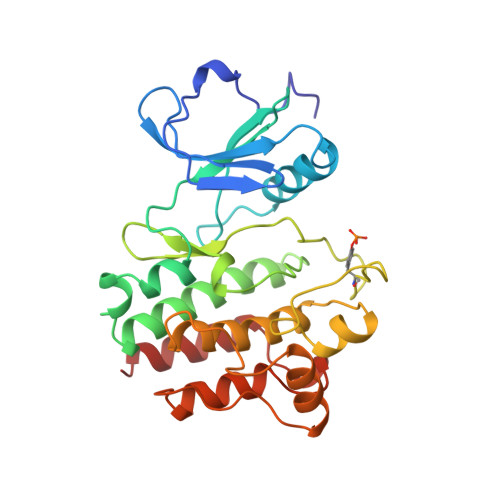
Structural basis for activation of human lymphocyte kinase Lck upon tyrosine phosphorylation.
Yamaguchi, H., Hendrickson, W.A.(1996) Nature 384: 484-489
- PubMed: 8945479 Search on PubMed
- DOI: https://doi.org/10.1038/384484a0
- Primary Citation Related Structures:
3LCK - PubMed Abstract:
Regulation through phosphorylation is a characteristic of signalling pathways and the lymphocyte kinase Lck (p56lck) both performs phosphorylation and is affected by it. Lck is a Src-family tyrosine kinase expressed in T lymphocytes, where it participates in the cellular immune response. Like all Src homologues, it comprises SH3, SH2 and kinase domains. Lck associates through its distinctive amino-terminal segment with the cytoplasmic tails of either T-cell co-receptor, CD4 or CD8-alpha. Activated Lck phosphorylates T-cell receptor zeta-chains, which then recruit the ZAP70 kinase to promote T-cell activation. Lck is activated by autophosphorylation at Tyr 394 in the activation loop and it is inactive when Tyr 505 near the carboxy terminus is phosphorylated and interacts with its own SH2 domain. Here we report the crystal structure of the Lck tyrosine kinase domain (LCKK) in its activated state at 1.7 A resolution. The structure reveals how a phosphoryl group at Tyr 394 generates a competent active site. Comparisons with other kinase structures indicate that tyrosine phophophorylation and ligand binding may in general elicit two distinct hinge-like movements between the kinase subdomains. From modelling studies, we suggest a basis for inhibition by phosphorylation at Tyr 505.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York 10032, USA.
Organizational Affiliation: