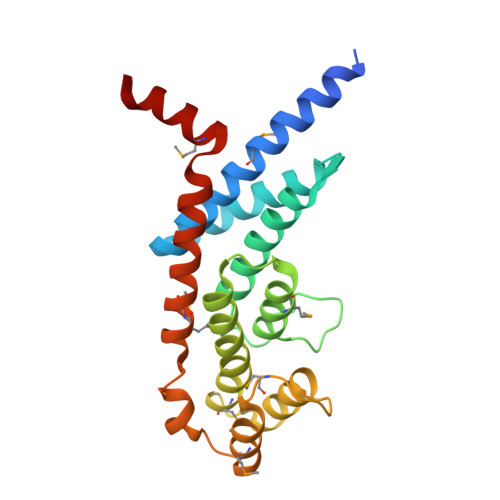
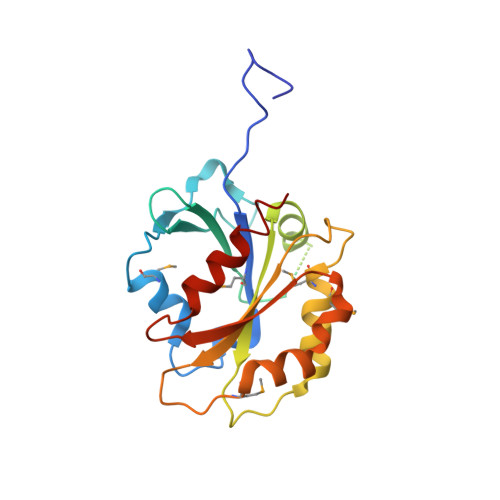
Structural Basis of Selective Ubiquitination of TRF1 by SCF(Fbx4)
Zeng, Z., Wang, W., Yang, Y., Chen, Y., Yang, X., Diehl, J.A., Liu, X., Lei, M.(2010) Dev Cell 18: 214-225
- PubMed: 20159592 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.devcel.2010.01.007
- Primary Citation Related Structures:
3L82 - PubMed Abstract:
TRF1 is a critical regulator of telomere length. As such, TRF1 levels are regulated by ubiquitin-dependent proteolysis via an SCF E3 ligase where Fbx4 contributes to substrate specification. Here, we report the crystal structure of the Fbx4-TRF1 complex at 2.4 A resolution. Fbx4 contains an unusual substrate-binding domain that adopts a small GTPase fold. Strikingly, this atypical GTPase domain of Fbx4 binds to a globular domain of TRF1 through an intermolecular beta sheet, instead of recognizing short peptides/degrons as often seen in other F-box protein-substrate complexes. Importantly, mutations in this interface abrogate Fbx4-dependent TRF1 binding and ubiquitination. Furthermore, the data demonstrate that recognition of TRF1 by SCF(Fbx4) is regulated by another telomere protein, TIN2. Our results reveal an atypical small GTPase domain within Fbx4 as a substrate-binding motif for SCF(Fbx4) and uncover a mechanism for selective ubiquitination and degradation of TRF1 in telomere homeostasis control.
- Howard Hughes Medical Institute, University of Michigan Medical School, 1150 West Medical Center Drive, Ann Arbor, MI 48109, USA.
Organizational Affiliation: