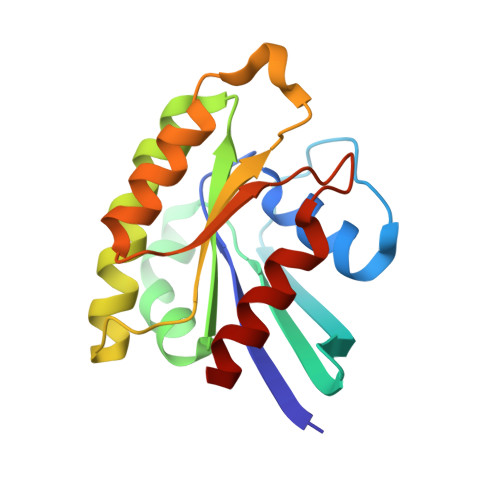
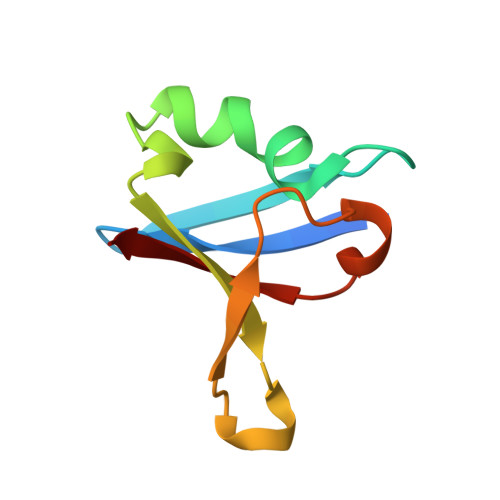
What makes Ras an efficient molecular switch: a computational, biophysical, and structural study of Ras-GDP interactions with mutants of Raf.
Filchtinski, D., Sharabi, O., Ruppel, A., Vetter, I.R., Herrmann, C., Shifman, J.M.(2010) J Mol Biology 399: 422-435
- PubMed: 20361980 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.03.046
- Primary Citation Related Structures:
3KUC, 3KUD - PubMed Abstract:
Ras is a small GTP-binding protein that is an essential molecular switch for a wide variety of signaling pathways including the control of cell proliferation, cell cycle progression and apoptosis. In the GTP-bound state, Ras can interact with its effectors, triggering various signaling cascades in the cell. In the GDP-bound state, Ras looses its ability to bind to known effectors. The interaction of the GTP-bound Ras (Ras(GTP)) with its effectors has been studied intensively. However, very little is known about the much weaker interaction between the GDP-bound Ras (Ras(GDP)) and Ras effectors. We investigated the factors underlying the nucleotide-dependent differences in Ras interactions with one of its effectors, Raf kinase. Using computational protein design, we generated mutants of the Ras-binding domain of Raf kinase (Raf) that stabilize the complex with Ras(GDP). Most of our designed mutations narrow the gap between the affinity of Raf for Ras(GTP) and Ras(GDP), producing the desired shift in binding specificity towards Ras(GDP). A combination of our best designed mutation, N71R, with another mutation, A85K, yielded a Raf mutant with a 100-fold improvement in affinity towards Ras(GDP). The Raf A85K and Raf N71R/A85K mutants were used to obtain the first high-resolution structures of Ras(GDP) bound to its effector. Surprisingly, these structures reveal that the loop on Ras previously termed the switch I region in the Ras(GDP).Raf mutant complex is found in a conformation similar to that of Ras(GTP) and not Ras(GDP). Moreover, the structures indicate an increased mobility of the switch I region. This greater flexibility compared to the same loop in Ras(GTP) is likely to explain the natural low affinity of Raf and other Ras effectors to Ras(GDP). Our findings demonstrate that an accurate balance between a rigid, high-affinity conformation and conformational flexibility is required to create an efficient and stringent molecular switch.
- Physikalische Chemie I, Fakultät für Chemie und Biochemie, Ruhr-Universität-Bochum, Universitätstr. 150, 44780 Bochum, Germany.
Organizational Affiliation: